Ever year, clinicians perform over 3.6 billion diagnostic x-ray examinations. Currently, trained radiologists interpret these billions of scans, but there is significant interest in developing automated systems for x-ray interpretation to assist radiologists and augment radiology expertise in underserved areas. This post will describe some recent data sets and methods applied to the problem of chest x-ray interpretation. (As a bonus, the final section also discusses the highly scientific “banana equivalent radiation dose.”)
The Goal
The goal of automated chest x-ray interpretation is to identify and localize abnormal findings in chest x-rays, such as pneumonia, tumors, and hernias. In the future, automated systems will likely streamline the radiology workflow by providing an initial x-ray interpretation to radiologists, and by prioritizing which scans should be read first (e.g. because they show worrisome pathology.)
Here are a few reasons why chest x-ray interpretation is a good medical imaging problem to solve:
- Billions of chest x-rays are collected each year, so building large data sets is feasible. Also, any high-performing automated method would be practically useful for many health systems.
- As a side bonus, chest x-rays are 2D, which makes them computationally easier to handle than 3D radiology studies like CT or MRI.
How Automated Chest X-Ray Interpretation Works
The majority of automated chest x-ray interpretation models described in the literature are convolutional neural networks (CNNs). CNNs are described in detail in this post.
In brief, CNNs process images hierarchically. First, the CNN identifies simple characteristics of the image, like edges. Then the CNN combines these simple characteristics to detect more complicated characteristics, like clouded areas of the lungs indicating pneumonia.
Chest X-Ray Data Set Format
Chest x-ray data are needed in order to train automated chest x-ray interpretation models. The chest x-ray data sets consist of images paired with labels. One example in a chest x-ray data set looks like this:
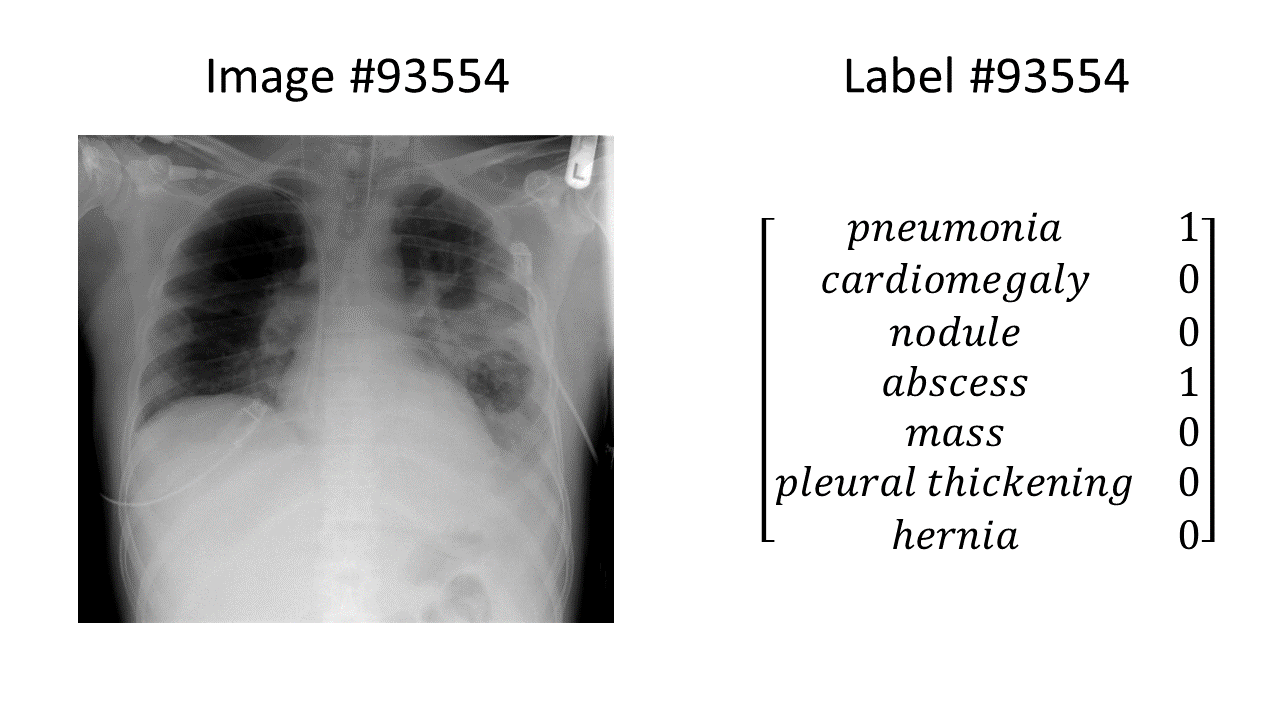
(the chest x-ray in the above example is from here)
In this example, we have a chest x-ray showing pneumonia and an abscess, and a binary label vector indicating that pneumonia = 1 (present), abscess = 1 (present), and all the other findings of interest = 0 (absent). Every example in the entire data set is formatted in the same way: an image paired with a binary label vector, for the same fixed set of labels in the same order.
Three important considerations are the data set size, the number of labels available, and the quality of labels available:
- Data Set Size: bigger is better! We want as many labeled images as possible.
- Number of labels: more is better! Because the models are trained using the labels, the only diseases the model can learn to detect are the diseases denoted in the labels. If there are three diseases of interest, then every single image in the data set must be labeled with “yes or no” for each of those three diseases of interest. If a certain disease is not included – for example if “pneumonia” were left out – then any supervised model trained on the data set would not be able to learn to detect pneumonia.
- Quality of labels: we want the labels to be as accurate as possible.
Where do the labels come from?
Often, the labels (which diseases are present in which image) are obtained using artificial intelligence approaches applied to the free-text note that was written by a past radiologist to describe the image. A note might look something like this: “There is pneumonia in the right upper lobe. The heart is enlarged. There is no pneumothorax, nodules, or masses.”
Getting the labels out of a note is useful because it means a person doesn’t have to write down all the labels by hand. However, it presents a problem if the automated method for obtaining labels is inaccurate (more on that below).
Real Chest X-Ray Data Sets and Models!
Here are some examples of publicly-available chest x-ray data sets, and a few accompanying models. Some papers publish both a new chest x-ray data set and a new model. Other papers publish only a new model, using an existing data set.
Data Set: ChestX-ray8 (May 5, 2017)
The ChestX-ray8 data set is described in this paper. It was created by the National Institutes of Health (NIH) Clinical Center, and includes 108,948 chest x-rays from 32,717 unique patients.
Each chest x-ray is labeled with eight disease labels: atelectasis (collapsed lung tissue), cardiomegaly (enlarged heart), effusion (excess liquid), infiltration, mass, nodule, pneumonia, and pneumothorax (air in the chest.) Thus, supervised models trained on this data set can be trained to detect these 8 diseases.
Data Set: ChestX-ray14. Model: CheXNet (November 14, 2017).
ChestX-ray14 is built off of ChestX-ray8, but it contains 14 disease labels instead of 8. ChestX-ray14 adds the following disease labels on top of ChestXray-8’s labels: consolidation (a region of lung tissue that is full of liquid instead of air), hernia, edema, emphysema (e.g. from smoking), fibrosis (hardening/scarring of the lungs), and “pleural thickening” (a lung disease where scarring makes the lining of the lungs thicker.)
CheXNet is a model for automated chest x-ray interpretation that was trained using the ChestX-ray14 data set. The model is a 121-layer convolutional neural network that can also make “heat maps” highlighting areas of the chest x-ray that were important for the model’s final decision.
Here’s a cartoon example of a yellow “heat map” highlighting the region of pneumonia/abscess from the chest x-ray example shown previously:

The authors claim that the CheXNet model achieves “radiologist-level performance.” However, not everyone agrees with that statement. The radiologist Luke Oakden-Rayner has an interesting blog post in which he describes several problems with the ChestX-ray14 data set, including problems with the labeling accuracy and whether the labels reflect what’s truly in the images. The general philosophical point is that because deep learning is flexible and powerful, a deep learning model is capable of learning incorrect diagnoses for chest x-rays if you train the model using incorrect diagnosis labels (“garbage in, garbage out”). In order to develop machine learning systems that have real-world applicability, we need to make sure that the labels we use to train the models truly reflect what diseases are present in the corresponding chest x-ray images.
Data Set: CheXpert (January 21, 2019)
CheXpert and MIMIC-CXR are the newest and biggest chest x-ray data sets, both released this past January.
CheXpert contains 224,316 scans from 65,240 patients, which is about twice as many scans and twice as many patients as ChestX-ray8. It includes these 14 labels (which are not exactly the same as the 14 ChestX-ray14 labels):
- cardiomegaly
- “enlarged cardiomegaly”: I’m honestly not sure what this is, since cardiomegaly is by definition an enlarged heart, so “enlarged cardiomegaly” just means “enlarged enlarged heart.” Perhaps they mean “enlarged cardiac silhouette” which can be caused by an enlarged heart OR by other problems, like fluid in the sac that surrounds the heart.
- lung lesion: this is a broad category, as a “lesion” can be caused by many disease processes
- lung opacity: this is also a broad category – basically “white blob in the lungs”
- edema
- consolidation
- pneumonia
- atelectasis
- pneumothorax
- pleural effusion
- “pleural other”: this seems like a catch-all category for other problems with the sacks around the lungs
- fracture: for example, a rib fracture
- support devices
- no finding
Dr. Rayner also has an opinion about the CheXpert data set. Tldr: CheXpert is a better data set than ChestXray14 because it’s bigger and it includes better-quality labels.
The CheXpert paper describes the results of a convolutional neural network model trained on the CheXpert data set. Similar to the CheXNet model described in the previous section, the CheXpert model also produces “heat maps” highlighting important areas of the images.
Data Set: MIMIC-CXR (January 21, 2019)
Last but not least…MIMIC-CXR was released this year on exactly the same day as the CheXpert data set (coincidence? I think not!) The data set is available through the MIMIC-CXR Database web page.
MIMIC-CXR uses the same labeling system as CheXpert, so it includes the same 14 CheXpert labels listed above on all of its images. MIMIC-CXR includes 371,920 chest x-rays from 227,943 imaging studies.
Takeaways
Since 2017, over 700,000 labeled chest x-ray images have been released to the public to accelerate development of machine learning models for automated chest x-ray interpretation. It is an exciting time to work in the automated medical imaging interpretation field!
About the Featured Image
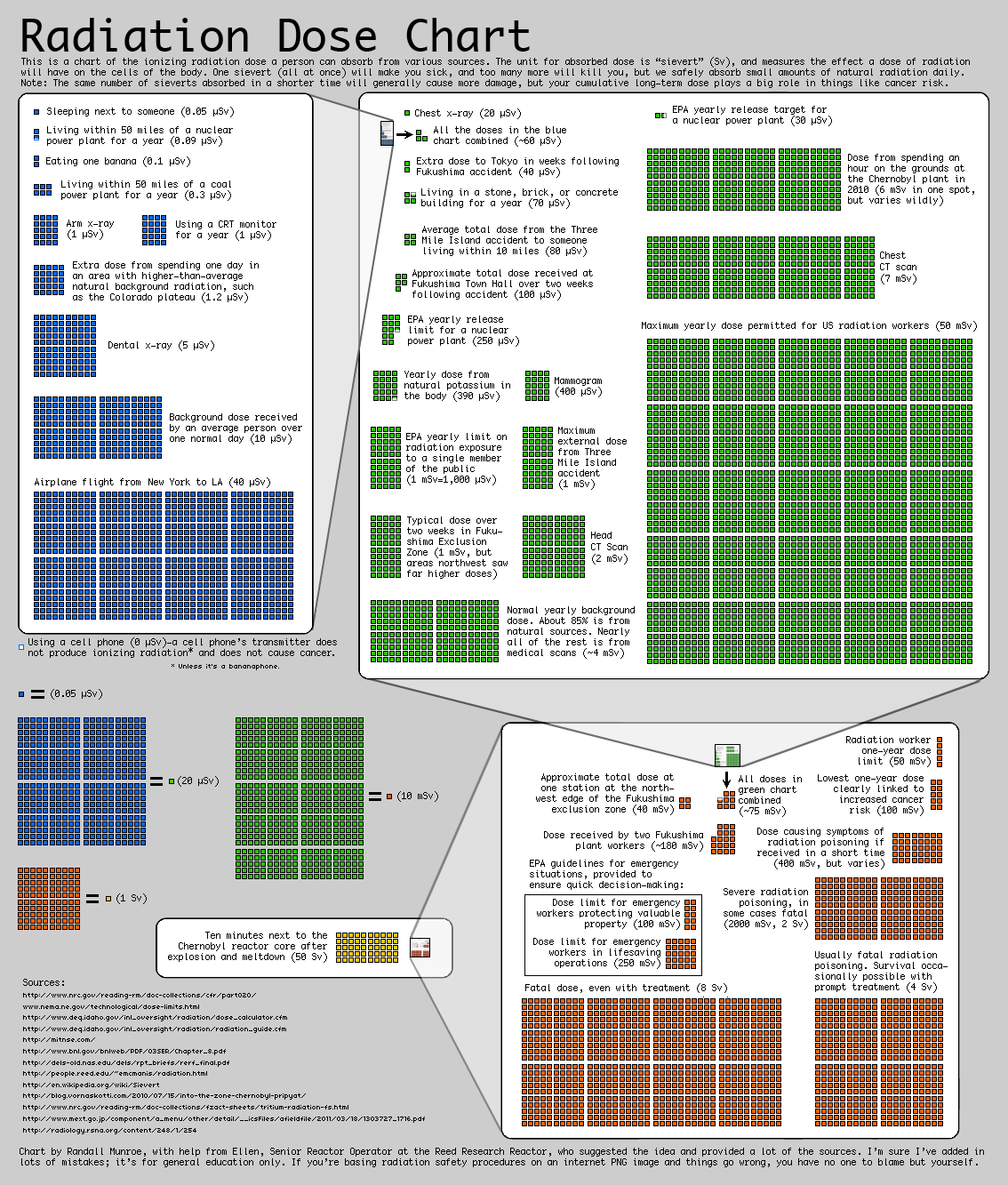
What’s going on with the banana smiley face in the featured image? (banana image source). I learned today about “banana equivalent dose” which is an informal indicator of ionizing radiation exposure, based on the radiation dose from eating one banana. Bananas contain a lot of potassium, including the radioactive isotope potassium-40. But, never fear: eating one banana is a minuscule dose of radiation, far less than the amount of radiation you’re exposed to just by being alive on Earth for one day.
Here is a fun chart on radiation doses from https://xkcd.com/radiation/ that includes “eating one banana” as two blue boxes in the upper right-hand corner:

Want to be the first to hear about my articles bridging healthcare, artificial intelligence, and business—and get a free list of my favorite health AI resources? Sign up here.


Comments are closed.