Classification labels specifying the presence and absence of abnormalities are necessary to train computer vision models on radiology images. However, obtaining these classification labels by hand is time-consuming and limits the size of the final dataset as well as the number of abnormalities that can be considered. In this post we’ll overview an easily customizable technique, SARLE, for extracting structured abnormality and location labels automatically from the free-text reports that accompany each radiology image in a hospital database.
What are radiology reports?
Every time a patient is imaged, a radiologist interprets the image and writes a report summarizing the normal and abnormal findings. An excerpt from a chest x-ray report might read, “There is a nodule in the right lung. The left lung is clear. There is cardiomegaly without pericardial effusion.”
What is radiology label extraction?
Radiology label extraction is the process of obtaining binary abnormality labels from a free text radiology report. It is more complex than merely identifying if an abnormality is mentioned because radiologists will often specifically note abnormalities that are absent or those that have resolved relative to a previous image. For example, “Groundglass opacities have resolved”, “The chest tube has been removed”, or “Left upper lobe nodule is no longer appreciated”.
Why radiology label extraction is difficult
Radiology label extraction is difficult for several reasons:
(1) as previously mentioned, some abnormalities are included in the report because they are present, while others are noted because they are absent, meaning negation/normality detection is needed;
(2) there are hundreds of possible abnormalities and many of them have synonyms or several different ways of being described (e.g., enlarged heart==cardiomegaly, pleural effusion==pleural fluid accumulation==fluid in the pleural space),
(3) there are many descriptive modifier terms (e.g., indicating texture, general size, measured size, size relative to a previous image, severity, and so on) and some of these descriptors may be the difference between something being normal or abnormal. For example, “lymphadenopathy” or enlarged lymph nodes are determined by a size greater than 1 centimeter. Lymphadenopathy may be described as “lymphadenopathy,” “adenopathy,” “enlarged lymph nodes,” or simply with a measurement, “1.7 cm mediastinal lymph node” or “22 mm lymph node in the right axilla.” Sometimes borderline enlarged lymph nodes are mentioned (e.g., “9 mm lymph node”) and other times lymph nodes that used to be large but are now normal are mentioned (e.g., “formerly 1.1 cm lymph node now measures 0.3 cm”).
(4) if you are interested in the anatomical location of abnormalities, there are also synonyms for certain anatomical locations (e.g., “left upper lobe==left superior lobe”) and some locations are only implied by the nature of the abnormality (e.g., pneumonia is a lung infection by definition, cardiomegaly is an enlarged heart by definition).
Why radiology label extraction is easier than dealing with other kinds of natural language
On the other hand, there are also some aspects of radiology label extraction that make it easier than general summarization tasks on natural language. Radiology notes are focused on a relatively narrow topic, which limits the kinds of phrases that are used (relative to, say, a novel or a book of poetry) and radiology notes have generally good grammar, spelling, and sentence structure.
Why radiology label extraction is useful in computer vision
Computer vision models perform better with more data. They also require structured labels for training, whether that’s a binary vector of presence/absence labels to train a classification model or a pixel-level tracing (segmentation map) to train a segmentation model. Let’s consider the easiest type of labels to obtain: classification labels. If we want to build a classifier dataset that includes 50 different abnormalities across 100,000 chest x-rays, we would need a radiologist to manually record 5,000,000 labels. At one second per label, that’s 1,388 hours which works out to 173 eight-hour days of really, really tedious work–to produce a single dataset for one imaging modality. There is thus a lot of interest in automated radiology label extraction, which produces exactly the classification labels we need from the existing free-text reports automatically with far less manual work.
Approaches to radiology label extraction
There are different ways to categorize label extraction approaches:
Method: The two major categories of method are rule-based and machine-learning-based. I remember scoffing at the idea that anyone would ever use rules in this age of massive language neural nets but it turns out that rules can work excellently for radiology label extraction because radiology notes are well-organized and focused on a limited topic.
Input data: the input to the method can be a whole note, a whole sentence, or a phrase.
Labels considered: the method may consider only one abnormality label, a few, or many. The method may not even consider abnormality labels at all–it may be focused on only anatomical locations, for example. Or, the method may consider both abnormalities and locations (e.g. SARLE).
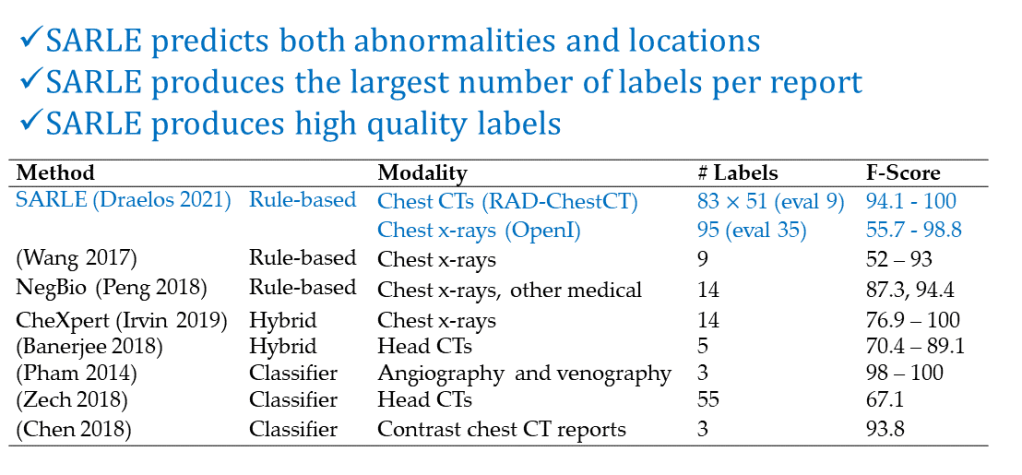
Several examples of radiology label extraction methods are summarized in Appendix Table B2 of this paper.
SARLE: Sentence Analysis for Radiology Label Extraction
SARLE is a publicly available, high-performance, easily customizable Python framework for radiology label extraction. It’s fast, easy to use, easy to adapt to a new type of radiology report or new set of labels, and has minimal dependencies. The core logic is fewer than 300 lines of Python code. It’s also the only radiology label extraction framework (as far as I’m aware) that extracts both abnormalities and locations: for each abnormality a corresponding location is provided, meaning the label output is actually a location x abnormally matrix rather than an abnormality vector.
SARLE has two steps:
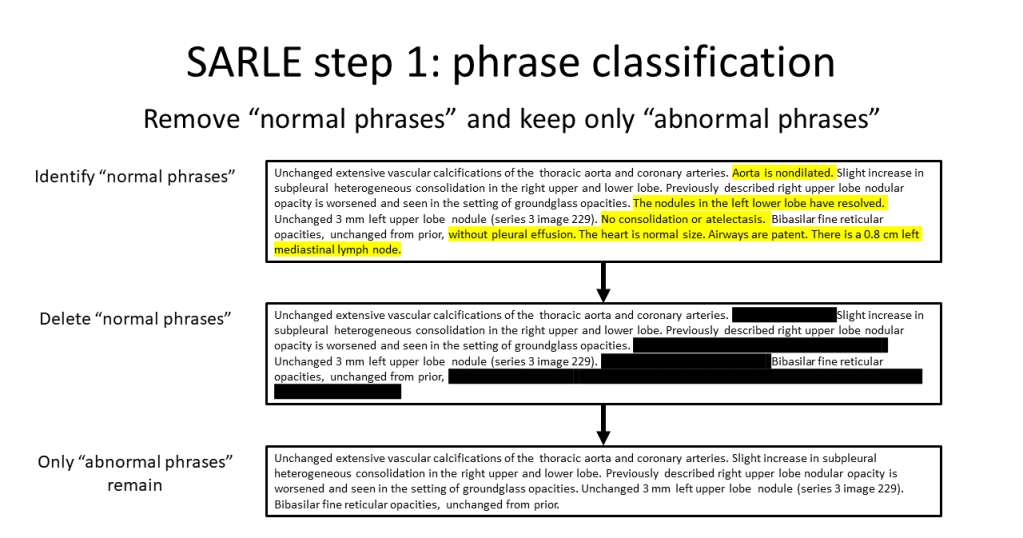
In the first step, a sentence classifier distinguishes between normal sentences (describing normal findings or lack of abnormalities) and abnormal sentences (describing presence of abnormal findings). All normal sentences are then discarded. There are two variants of SARLE: in SARLE-Hybrid, a machine learning classifier performs sentence classification, and in SARLE-Rules, a rule-based method performs sentence classification.
The second step is a term search. All the abnormal sentences are fed into a term search that uses medical synonyms to identify mentions of abnormalities and anatomical locations. Because only abnormal sentences remain, any mention of an abnormality indicates that it is present.

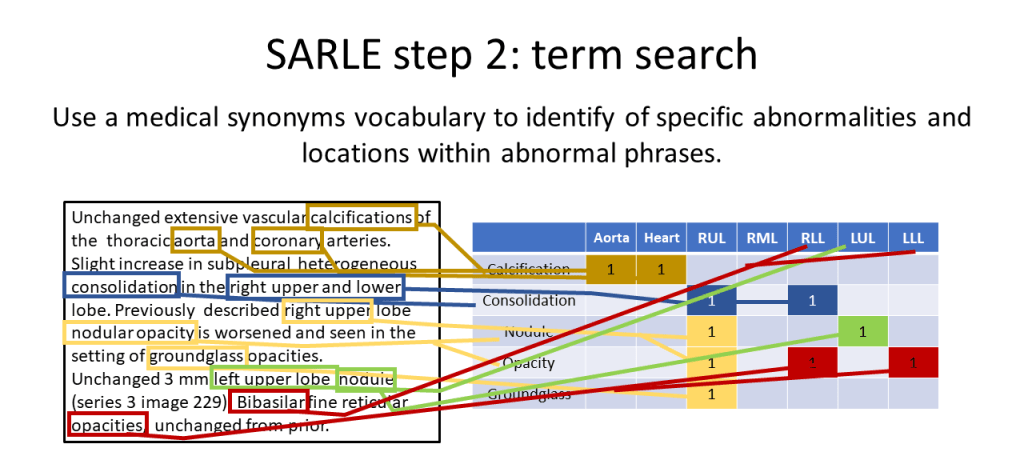
The 83 abnormalities SARLE extracts from radiology reports are shown in the table below:
| Lung (22) | airspace disease, air trapping, aspiration, atelectasis, bronchial wall thickening, bronchiectasis, bronchiolectasis, bronchiolitis, bronchitis, consolidation, emphysema, hemothorax, interstitial lung disease, lung resection, mucous plugging, pleural effusion, infiltrate, pleural thickening, pneumonia, pneumonitis, pneumothorax, pulmonary edema, scattered nodules, septal thickening, tuberculosis |
| Lung Patterns (5) | bandlike or linear, groundglass, honeycombing, reticulation, tree in bud |
| Additional (47) | arthritis, atherosclerosis, aneurysm, breast implant, breast surgery, calcification, cancer, catheter or port, cavitation, clip, congestion, cyst, debris, deformity, density, dilation or ectasia, distention, fibrosis, fracture, granuloma, hardware, hernia, infection, inflammation, lesion, lucency, lymphadenopathy, mass, nodule, nodule > 1 cm, opacity, plaque, postsurgical, scarring, scattered calcifications, secretion, soft tissue, staple, stent, suture, transplant, chest tube, tracheal tube, GI tube (includes NG and GJ tubes) |
| Heart (9) | cabg (coronary artery bypass graft), cardiomegaly, coronary artery disease, heart failure, heart valve replacement, pacemaker or defibrillator, pericardial effusion, pericardial thickening, sternotomy |
The 51 locations SARLE extracts are:
- Lungs: left upper lobe, lingula, left lower lobe, right upper lobe, right middle lobe, right lower lobe, right lung, left lung, lung, interstitial, centrilobular, subpleural, airways.
- Heart: heart, mitral valve, aortic valve, tricuspid valve, pulmonary valve.
- Great vessels: aorta, superior vena cava, inferior vena cava, pulmonary artery, pulmonary vein.
- General: right, left, anterior, posterior, superior, inferior, medial, lateral.
- Abdomen: abdomen, esophagus, stomach, intestine, liver, gallbladder, kidney, adrenal gland, spleen, pancreas.
- Other: thyroid, breast, axilla, chest wall, rib, spine, bone, mediastinum, diaphragm, hilum.
Performance
SARLE’s performance was analyzed for 427 chest CT reports across 9 labels with manually obtained ground truth. SARLE achieves high performance as shown in the table below:
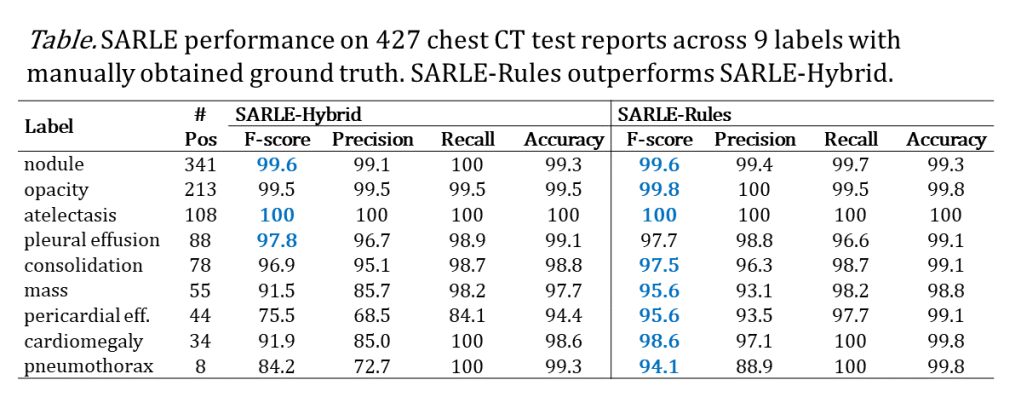
Overall, SARLE-Rules (which uses rules for sentence classification) outperformed SARLE-Hybrid (which uses machine learning for sentence classification). This is most likely because SARLE-Rules is technically performing phrase-level classification – i.e. it’s able to identify sub-parts of sentences that are normal or abnormal, as opposed to SARLE-Hybrid which is implemented at the whole sentence level. That works particularly well for sentences mentioning both a normal finding and an abnormal finding, e.g. “There is cardiomegaly without pericardial effusion.”
As a side note, the term search step of SARLE includes some sophisticated rules for handling abnormalities that depend on measurements, like the lymphadenopathy example mentioned before.
SARLE Code
SARLE code is publicly available here: https://github.com/rachellea/sarle-labeler
The code is structured so that it’s straightforward to adapt SARLE to your own dataset, your own abnormalities, and your own anatomical locations.
The script demo.py includes a demo of SARLE on real data and fake data:
- real data: SARLE is demonstrated on the OpenI dataset of chest x-ray reports.
- fake data: SARLE is demonstrated on some tiny handcrafted dataframes of fake data, to demonstrate the data format in a simple manner.
SARLE receives pandas dataframes as input. Details about their format is provided in the README and demo.py of the repository.
To customize the abnormalities SARLE detects, you can edit the vocabulary files directly:
- src/vocab/vocabulary_ct.py
- src/vocab/vocabulary_cxr.py
If you would like to customize SARLE’s locations, you can do so in
- src/vocab/vocabulary_locations.py
Paper
For more details about SARLE, or to cite SARLE, you can check out this paper: “Machine-Learning-Based Multiple Abnormality Prediction with Large-Scale Chest Computed Tomography Volumes.”
Summary
SARLE is a high-performance framework for radiology label extraction, publicly available as Python code with minimal dependencies. It’s easy to adapt to new datasets and to customize it for your own list of abnormalities and locations of interest. If you have any questions about deploying SARLE on your own dataset, feel free to reach out to me via this Contact page!
About the Featured Image
The featured image is composed of this paper image from Wikipedia (CC BY-SA 3.0) and this chest x-ray image from Wikipedia (Creative Commons 1.0).
Want to be the first to hear about my articles bridging healthcare, artificial intelligence, and business—and get a free list of my favorite health AI resources? Sign up here.

Comments are closed.