This post provides an overview of chest CT scan machine learning organized by clinical goal, data representation, task, and model.
A chest CT scan is a grayscale 3-dimensional medical image that depicts the chest, including the heart and lungs. CT scans are used for the diagnosis and monitoring of many different conditions including cancer, fractures, and infections.
Clinical Goal
The clinical goal refers to the medical abnormality that is the focus of the study. The following figure illustrates some example abnormalities, shown as 2D axial slices through the CT volume:
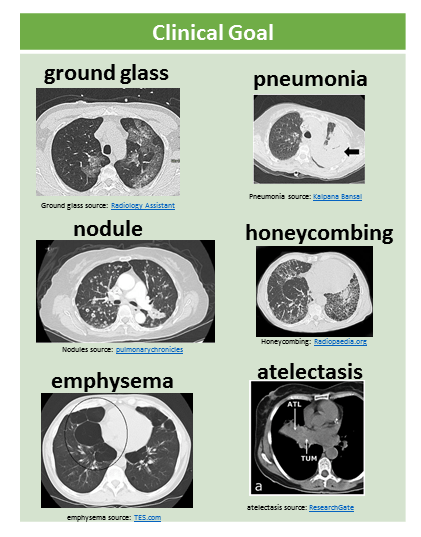
Image by Author. Sub-images from: ground glass Radiology Assistant, pneumonia Kalpana Bansal, nodules pulmonarychronicles, honeycombing Radiopaedia.org, emphysema TES.com, atelectasis ResearchGate
Many CT machine learning papers focus on lung nodules.
Other recent work has looked at pneumonia (lung infection), emphysema (a kind of lung damage that can be caused by smoking), lung cancer, or pneumothorax (air outside of the lungs rather than inside the lungs).
I have been focused on multiple abnormality prediction, in which the model predicts 83 different abnormal findings simultaneously.
Data
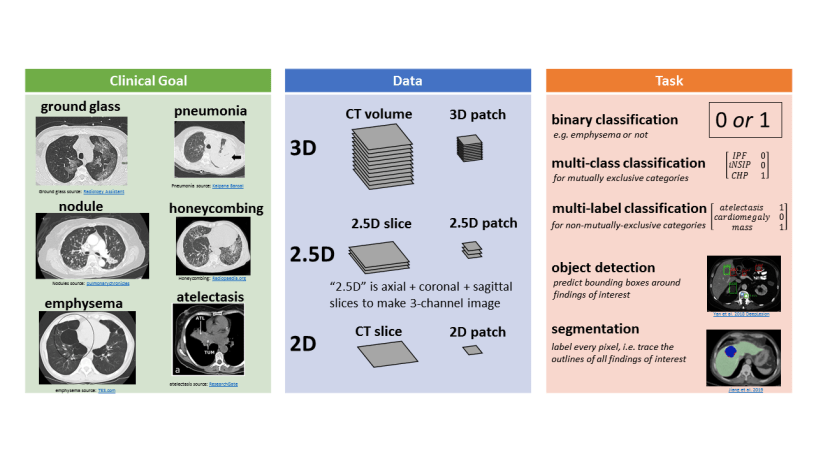
There are several different ways to represent CT data in a machine learning model, illustrated in this figure:

3D representations include a whole CT volume which is roughly 1000 x 512 x 512 pixels, and a 3D patch which can be large (e.g. half or a quarter of a whole volume) or small (e.g. 32 x 32 x 32 pixels).
2.5D representations make use of different perpendicular planes.
- The axial plane is horizontal like a belt, the coronal plane is vertical like a headband or old-style headphones, and the sagittal plane is vertical like the plane of a bow and arrow in front of an archer.
- If we take one axial slice, one sagittal slice, and one coronal slice, and stack them up into a 3-channel image, then we have a 2.5D slice representation.
- If this is done with small patches, e.g. 32 x 32 pixels, then we have a 2.5D patch representation.
Finally, 2D representations are also used. This could be a full slice (e.g. 512 x 512), or a 2D patch (e.g. 16 x 16, 32 x 32, 48 x 48). These 2D slices or patches are usually from the axial view.
Task
There are many different tasks in chest CT machine learning.
The following figure illustrates a few tasks:
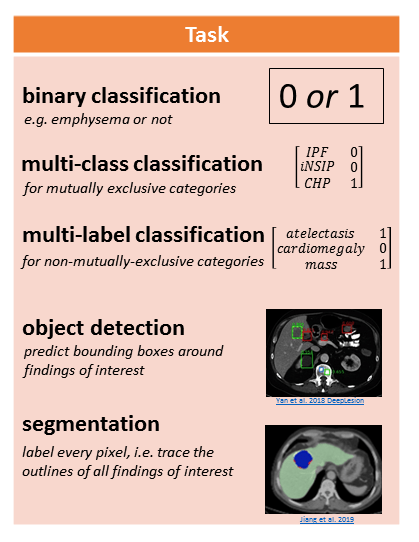

Image by Author. Sub-images from Yan et al. 2018 DeepLesion and Jiang et al. 2019
Binary classification involves assigning a 1 or 0 to the CT representation, for the presence (1) or absence (0) of an abnormality.
Multi-class classification is for mutually exclusive categories, like different clinical subtypes of interstitial lung disease. In this case the model assigns 0 to all categories except for 1 category.
Multi-label classification is for non-mutually-exclusive categories, like atelectasis (collapsed lung tissue), cardiomegaly (enlarged heart), and mass. A CT scan might have some, all, or none of these findings, and the model determines which ones if any are present.
Object detection involves predicting the coordinates of bounding boxes around abnormalities of interest.
Segmentation involves labeling every pixel, which is conceptually like “tracing the outlines of abnormalities and coloring them in.”
Different labels are needed to train these models. “Presence or absence” labels for abnormalities are needed to train classification models, e.g. [atelectasis=0, cardiomegaly = 1, mass = 0]. Bounding box labels are needed to train an object detection model. Segmentation masks (traced and filled in outlines) are needed to train a segmentation model. Only “presence or absence” labels are scalable to tens of thousands of CT scans, if these labels are extracted automatically from free-text radiology reports (e.g. the RAD-ChestCT data set of 36,316 CTs). Segmentation masks are the most time-consuming to obtain because they must be drawn manually on each slice; thus, segmentation studies typically use on the order of 100 – 1,000 CT scans.
Model
Convolutional neural networks are the most popular machine learning model used on CT data. For a 5-minute intro to CNNs, see this article.
- 3D CNNs are used for whole CT volumes or 3D patches
- 2D CNNs are used for 2.5D representations (3 channels, axial/coronal/sagittal), in the same way that 2D CNNs can take a 3-channel RGB image as input (3 channels, red/green/blue).
- 2D CNNs are used for 2D slices or 2D patches.
Some CNNs combine 2D and 3D convolutions. CNNs can also be “pretrained” which typically refers to first training the CNN on a natural image dataset like ImageNet and then refining the CNN’s weights on the CT data.
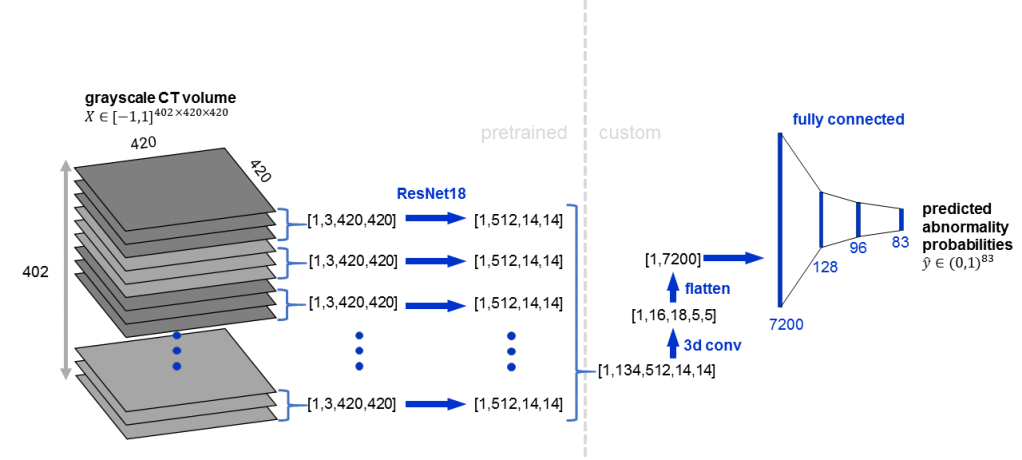
Here is an example architecture in which a pretrained 2D CNN (ResNet18) is applied to groups of 3 adjacent slices, followed by 3D convolution:
Interstitial Lung Disease Classification Examples
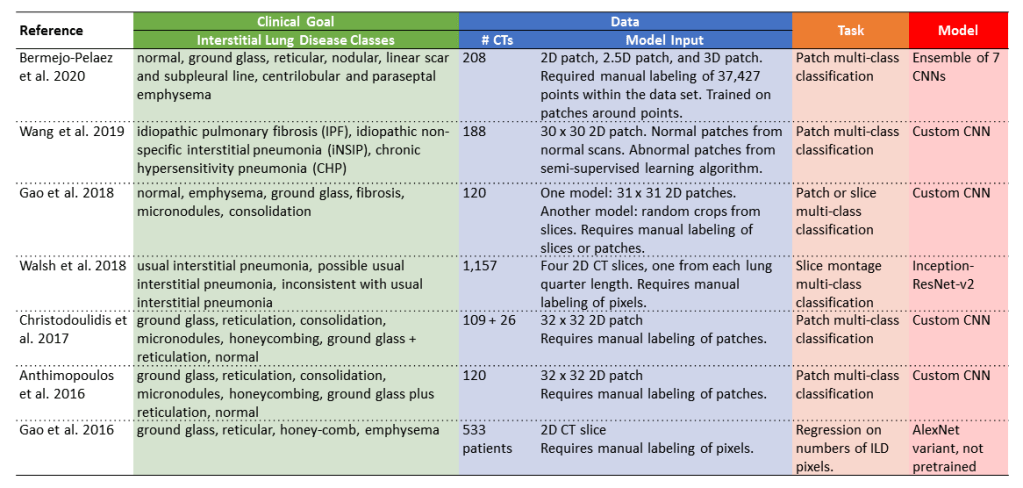
The following table includes several example studies focused on interstitial lung disease, organized by clinical goal, data, task, and model.
- Clinical goal: these papers are all focused on interstitial lung disease. The exact classes used differ between studies. Some studies focus on clinical groupings like idiopathic pulmonary fibrosis or idiopathic non-specific interstitial pneumonia (e.g. Wang et al. 2019 and Walsh et al. 2018). Other studies focus on lung patterns like reticulation or honeycombing (e.g. Anthimopoulos et al. 2016 and Gao et al. 2016).
- Data: the data sets consist of 100 – 1,200 CTs because all of these studies rely on manual labeling of patches, slices, or pixels, which is very time-consuming. The upside of doing patch, slice, or pixel-level classification is that it provides localization information in addition to diagnostic information.
- Task: the tasks are mostly multi-class classification, in which each patch or slice is assigned to exactly one class out of multiple possible classes.
- Model: some of the studies use custom CNN architectures, like Wang et al. 2019 and Gao et al. 2018, whereas other studies adapt existing CNN architectures like ResNet and AlexNet.
Additional Reading
- For a longer, more in-depth article on this topic, see Automatic Interpretation of Chest CT Scans with Machine Learning
- For an article about machine learning in chest x-rays, which are 2D medical images of the chest rather than 3D medical images of the chest, see Automated Chest X-Ray Interpretation
- For more info about CNNs, see Convolutional Neural Networks in 5 minutes and How Computers See: Intro to Convolutional Neural Networks
- For more details about segmentation tasks, see Segmentation: U-Net, Mask R-CNN, and Medical Applications
- For more details about classification tasks, see Multi-label vs. Multi-class Classification: Sigmoid vs. Softmax
Want to be the first to hear about my articles bridging healthcare, artificial intelligence, and business—and get a free list of my favorite health AI resources? Sign up here.

Comments are closed.