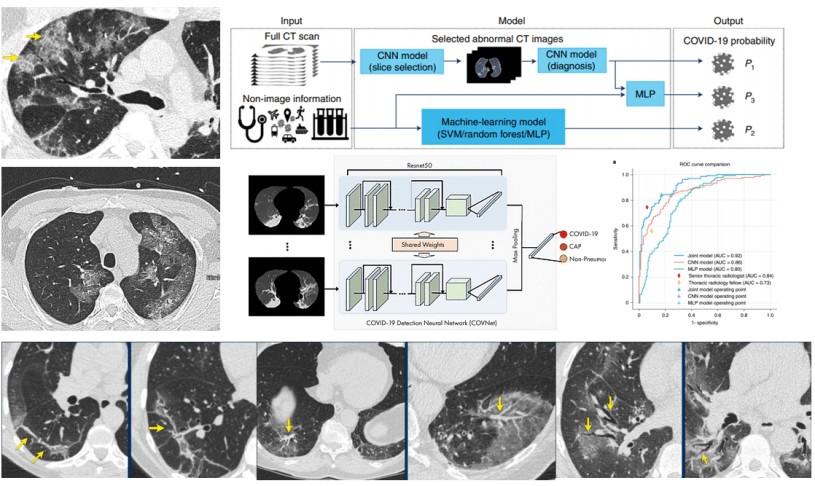
This post delves in to the use of CT scans in the COVID-19 pandemic, including current guidelines from medical experts (as of August 2020) and examples of recent research papers that use machine learning to make predictions from CT scans of COVID-19 patients.
Disclaimer: Nothing in this post is medical advice.
Diagnosis of COVID-19 with RT-PCR
The gold standard for diagnosis of COVID-19 is reverse transcription polymerase chain reaction (RT-PCR), which is a laboratory test that detects genetic material (RNA) from the COVID-19 virus:
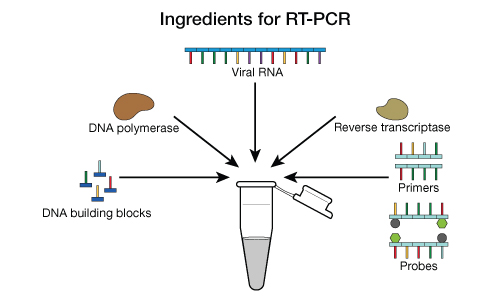
Figure Source: Mayo Clinic
The RT-PCR test is the “nasal swab test.” The nasal swab is intended to collect virus, if present, so that its RNA can be detected:
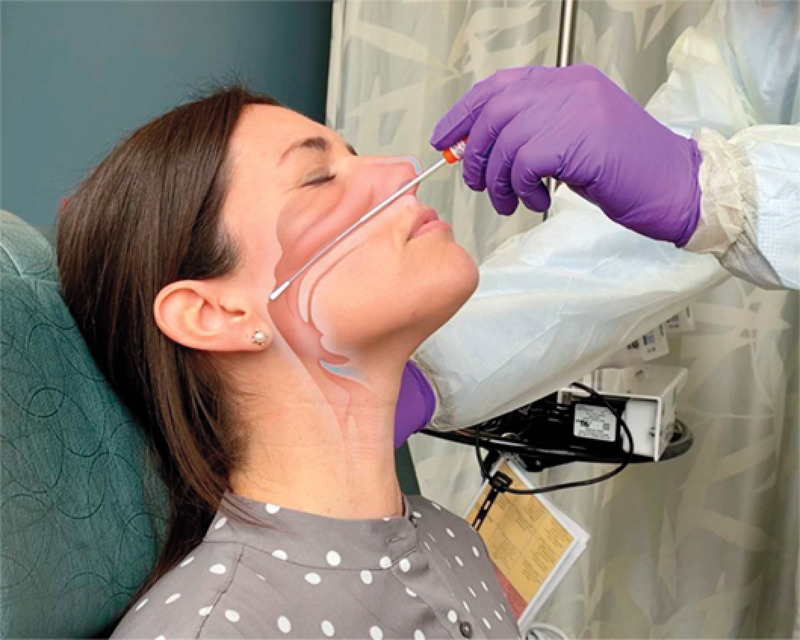
Figure Source: New England Journal of Medicine
The RT-PCR test has high specificity (true negative rate). The specificity is close to 100%, meaning that almost all healthy people are correctly identified as healthy.
The RT-PCR test has less sensitivity (aka true positive rate, aka recall), which measures the percentage of sick people correctly identified as sick. Sensitivity = (true positives) / (true positives + false negatives). This means that false negative tests can occur. For extra certainty that a patient is negative, multiple tests may be performed.
In spite of the imperfect sensitivity, RT-PCR is still the gold standard – the best test that we currently have for diagnosing COVID-19.
The Role of Chest CT in COVID-19
Given that RT-PCR is the gold standard for COVID-19 diagnosis, what is the role of chest CT in COVID-19?
The Fleischner Society published a multinational consensus statement in the journal Radiology in April 2020 that states (emphasis added):
Imaging is not indicated in patients suspected of having coronavirus disease 2019 (COVID-19) and mild clinical features unless they are at risk for disease progression.
Imaging is indicated in a patient with COVID-19 and worsening respiratory status.
In a resource-constrained environment, imaging is indicated for medical triage of patients suspected of having COVID-19 who present with moderate-to-severe clinical features and a high pretest probability of disease.
Some research papers have claimed that chest CT scans have high sensitivity or high specificity for COVID-19 diagnosis, and that therefore CT scans could potentially be used for diagnosis. However, a review by Raptis et al., “Chest CT and Coronavirus Disease (COVID-19): A Critical Review of the Literature to Date” identifies numerous methodological problems with these studies. Raptis et al. conclude,
Even in situations in which RT-PCR test results are negative, delayed, or not available, no data of which we are aware support CT as an adequate replacement test because its true sensitivity is unknown [and] CT findings lack specificity.
In summary, chest CT scans are NOT currently recommended for COVID-19 diagnosis or screening. However, chest CTs may be helpful for evaluating COVID-19 complications, triage in resource-constrained environments, or prediction of worsening vs. improvement:
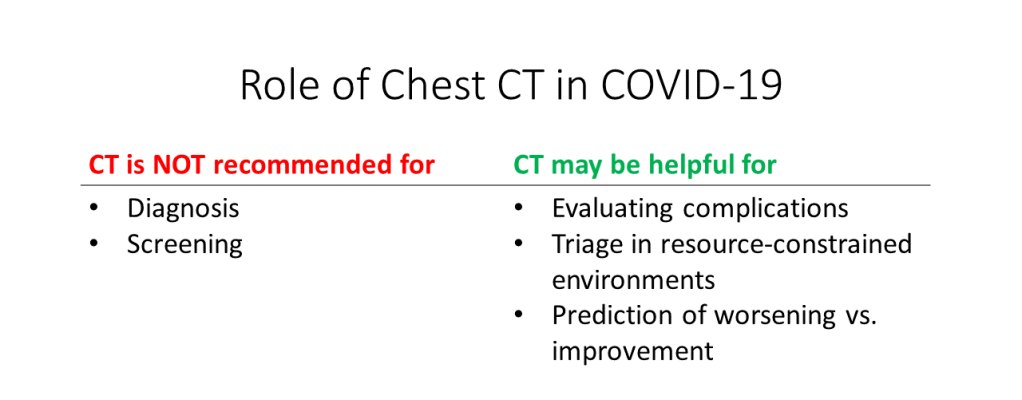

Figure by Author. Additional references: Radiopedia, Radiology Assistant
COVID-19 Appearance on Chest CT
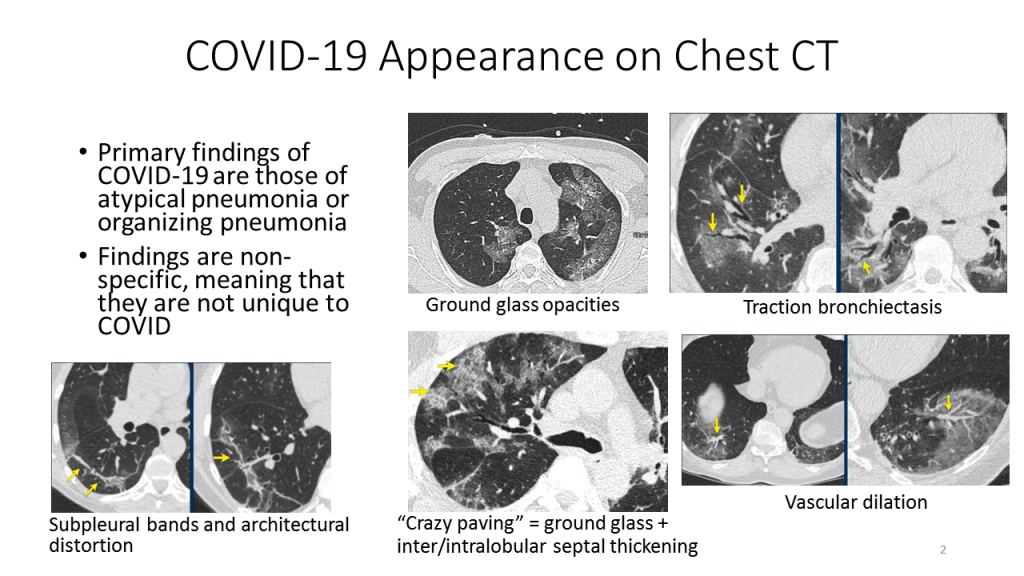
What does COVID-19 look like in a chest CT? The primary findings of COVID-19 are the findings of atypical pneumonia or organizing pneumonia. Such findings are non-specific, meaning that they are not unique to COVID-19 and can be seen in lung infections caused by bacteria or by other kinds of viruses.
The following figure, based on CT slices from Radiology Assistant, shows some of the findings that can be seen in COVID-19 patients on chest CT. These findings include ground glass opacities, “crazy paving”, traction bronchiectasis, vascular dilation, and architectural distortion:
Machine Learning in COVID-19 Chest CTs
There has been considerable interest in building machine learning models to help with the COVID-19 pandemic. In general, any medical machine learning models should only be deployed after extensive validation, in agreement with medical best practices, and under the guidance of medical professionals.
It is worth noting that the same model has the potential to be used in appropriate or inappropriate ways. For example, consider a COVID-19 diagnosis model, which is built to take in medical data and output the probability of COVID-19 diagnosis. This model could be used in line with medical guidelines to assist with triage in a resource-constrained environment. It could also be used in violation of medical guidelines to diagnose patients in place of the gold standard RT-PCR test.
All of the papers that I have seen so far about building machine learning models for chest CTs in COVID-19 are focused on development of models. This is distinct from deployment of models which requires a different kind of research focused on determining whether the model benefits clinicians and/or patients in a measurable way.
Now I will overview three papers that have built machine learning models on COVID-19 CT data. Each of the papers takes a different approach.
Background
All COVID-19 CT scan machine learning models are based on convolutional neural networks. If you are not familiar with convolutional neural networks, please read Convolutional Neural Networks (CNNs) in 5 minutes.
Chest CT scans are volumetric grayscale medical images that depict the heart and lungs. They are used in the diagnosis and management of a wide range of conditions including cancer, fractures, and infections. For more background on chest CT machine learning, please see Chest CT Scan Machine Learning in 5 minutes.
Paper #1

In this paper, the authors build and evaluate a model that takes a stack of CT slices as input, and predicts whether the scan shows COVID-19 pneumonia, community acquired pneumonia (“CAP”), or no pneumonia.
All COVID-19 cases were confirmed positive using RT-PCR, to ensure that the ground truth was of good quality.
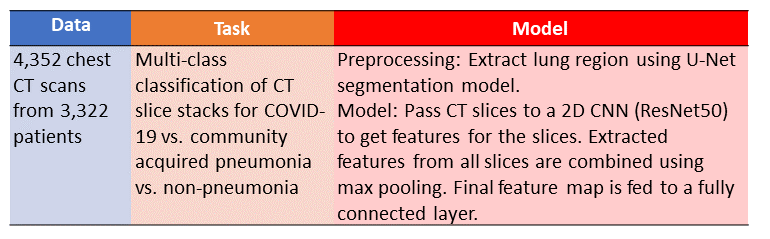
Table Summary of Li et al. paper. Image By Author
In this model, the CT scans are first preprocessed using a U-Net, which performs lung segmentation to extract out only the lung regions and exclude the heart and body wall.
Here is a diagram of the U-Net architecture:
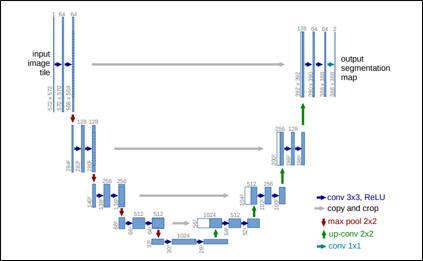
U-Net architecture, from Ronneberger et al. 2015
After this preprocessing step is complete, features are extracted from each CT slice using a ResNet50 convolutional neural network. The slice features are combined with max pooling and a final fully connected layer produces the predictions.
Here is an overall diagram of the “COVID-19 vs. CAP vs. non-pneumonia” model. Note that the U-Net preprocessing step is not explicitly shown:
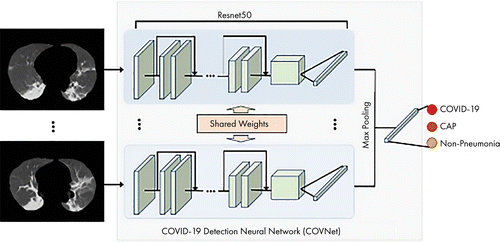
Model Architecture. From Li et al.
Results: The authors report good performance, with AUCs of 0.96, 0.95, and 0.98 for COVID-19, community-acquired pneumonia (CAP), and no pneumonia, respectively:
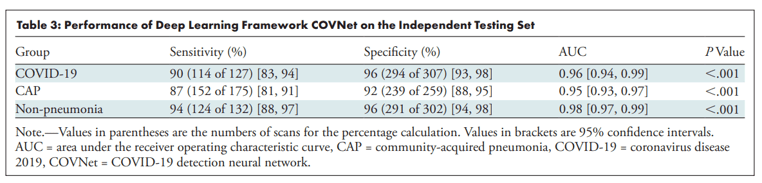
Model Results. From Li et al.
The authors also made Grad-CAM visualizations as a form of model explanation. Grad-CAM is a technique for creating a heatmap that shows where a model is focusing for a particular class. For more details on Grad-CAM see Grad-CAM: Visual Explanations from Deep Networks.
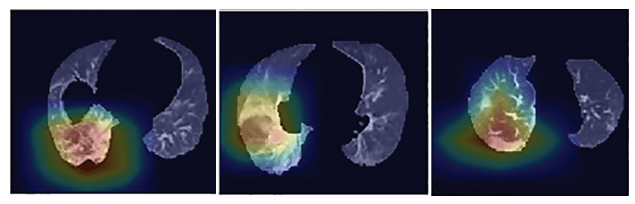
Grad-CAM visualizations. From Li et al.
The authors provide a good commentary on Grad-CAM in their discussion section:
a disadvantage of all deep learning methods is the lack of transparency and interpretability (eg, it is impossible to determine what imaging features are being used to determine the output). While we used a heatmap to visualize the important regions in the scans leading to the decision of the algorithm, heatmaps are still not sufficient to visualize what unique features are used by the model to distinguish between COVID-19 and CAP.
Overall, this was a clean research study, with a ground truth based on the RT-PCR gold standard, a well-explained model architecture, and high performance. The one aspect that would benefit from updating is the motivation, which the authors summarize as, “RT-PCR is considered the reference standard; however it has been reported that chest CT could be used as a reliable and rapid approach for screening of COVID-19.” Currently (as of August 2020), chest CT scans are not recommended for screening. A better motivation, more in line with current clinical recommendations, would be to use their high-performing CT scan model to help with triage in a resource-constrained environment.
Paper #2

The previous paper focused on a diagnosis model for COVID-19. Paper #2 by Huang et al. has a different goal: to investigate the relationship between the amount of lung opacification and COVID-19 severity.
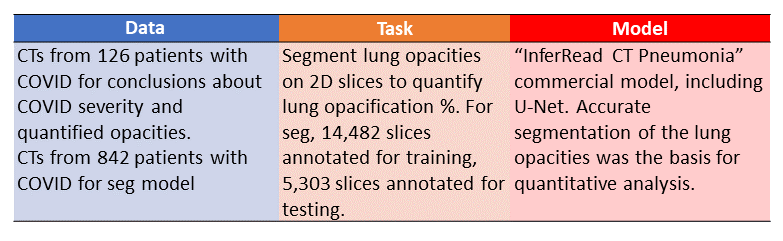

Table Summary of Huang et al study. Image by Author
On a chest CT scan the lungs are black because they are full of air. A “lung opacity” is a white splotch within the lungs, caused by material like water or pus accumulating inside of the lung tissue. Lung opacities are commonly seen in pneumonia, including pneumonia caused by COVID-19.
In order to quantify the amount of lung opacification, Huang et al. use a commercial machine learning model called “InferRead CT Pneumonia.” Because this commercial model is proprietary the authors do not provide a diagram of the architecture, but they do mention that it is based on a U-Net, the segmentation architecture described in the previous section.
The InferRead CT Pneumonia model traces the outlines of all the lung opacities (more specifically, it identifies all of the pixels that are part of a lung opacity):
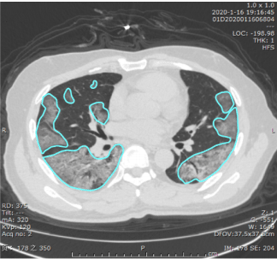
Example of lung opacities segmented by the deep learning model in Huang et al.
The percent of lung pixels that are opacified then serves as a quantitative measure of the extent of the lung opacification. A “100% opacified lung” would be all white instead of all black.
Huang et al. analyze 126 patients with differing severity of COVID-19 and investigate whether more clinically severe COVID-19 corresponds to quantitatively more lung opacification. COVID-19 diagnosis was confirmed with RT-PCR, and the patient’s clinical severity was measured using the “Diagnosis and Treatment Protocol of Novel Coronavirus” from the National Health Commission of China, which classifies patients into mild, moderate, severe, and critical:
- Mild type: patients have mild clinical symptoms without CT findings of pneumonia
- Moderate type: patients have fever and respiratory symptoms with CT findings of pneumonia
- Severe type: patients meet any of the following criteria: a) respiratory distress (respiratory rate ≥ 30 bpm) b) SpO2 ≤ 93% at rest c) PaO2/FiO2 ≤ 300 mmHg
- Critical type: patients meet any of the following criteria: a) respiratory failure with mechanical ventilation b) shock other organ dysfunction and ICU therapy.
Results: As you can see from the table below, the mild type patients had 0% lung opacification (i.e., fully healthy lungs), while the critical type patients had almost 50% lung opacification. A statistical test found that the difference in lung opacification across different clinical severity was significant with p < 0.001.
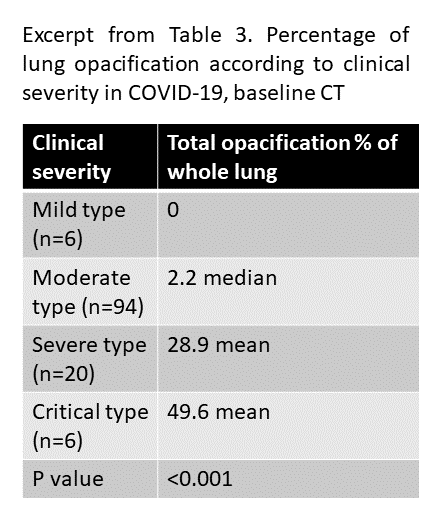
Image by Author, summarizing results from Huang et al.
Huang et al. conclude,
There were significant differences in lung opacification percentage, as measured by the deep learning algorithm, among patients with different clinical severity […] This automated tool for quantification of lung involvement may be used to monitor the disease progression and understand the temporal evolution of COVID-19.
Paper #3

Similar to paper #1, the goal of paper #3 by Mei et al. is to build a COVID-19 diagnosis model. However, while paper #1 uses only CT data as the input, paper #3 uses both CT data and clinical data as input.
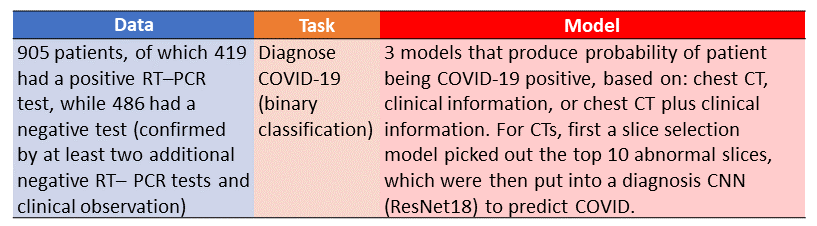

Table Summary of Mei et al. paper. Image by Author
Mei et al. summarize their study as follows:
In this study, we used artificial intelligence (AI) algorithms to integrate chest CT findings with clinical symptoms, exposure history and laboratory testing to rapidly diagnose patients who are positive for COVID-19.
The paper uses medical data from 905 patients, of which roughly half were positive for COVID-19. The task was binary classification, meaning patients are labeled with a one (COVID-19 positive) or a zero (COVID-19 negative).
The authors compare three different models: one model that uses only chest CT data, another model that uses only clinical information, and a third model that combines chest CT data and clinical information.
These three different models are summarized as different pathways through the figure below:
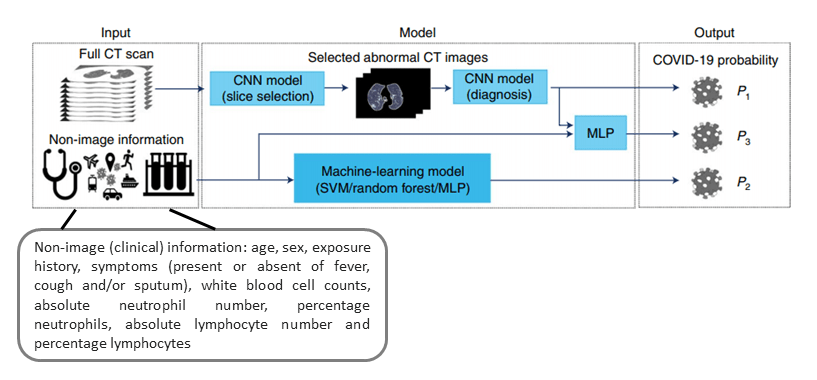
Model Diagram, modified from a figure in Mei et al.
The model that uses chest CT data involves two steps: first, a slice selection model picks out the top 10 most abnormal slices, and then these slices are fed into a “diagnosis CNN” (ResNet18 architecture) to predict COVID-19 status.
The model that combines chest CT data and clinical data involves feeding the output of the CT “diagnosis CNN” and the output of the clinical data model into a multilayer perceptron to produce the final prediction of COVID-19 status.
Results: The model combining CT and clinical data (joint model) had a 6% better AUROC than the model on CT data alone (P = 0.00146), and a 12% better AUROC than the model on clinical information alone (P < 1 × 10−4), showing that COVID-19 prediction was most effective when both CT data and other clinical data were used together. The authors also compared their algorithm to a senior thoracic radiologist, and found that “the algorithm performed equally well in sensitivity (P=0.05) in the diagnosis of COVID-19 as compared to a senior thoracic radiologist.”
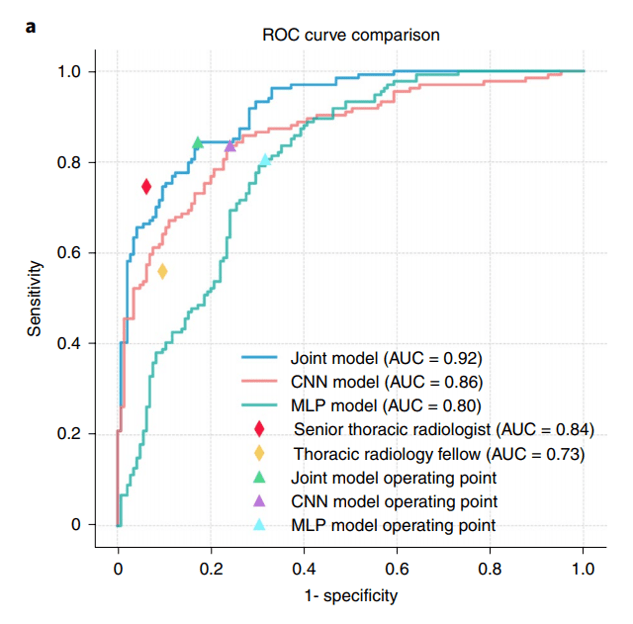
ROC curves from Mei et al.
Mei et al. conclude,
While chest CT is not as accurate as RT–PCR in detecting the virus, it may be a useful tool for triage in the period before definitive results are obtained.
Summary
- Chest CTs are volumetric grayscale medical images that depict the heart and lungs.
- 2D and 3D convolutional neural networks (CNNs) are applied to classify, box, or segment CT abnormalities.
- The gold standard for COVID-19 diagnosis is RT-PCR, not chest CT.
- In the context of COVID-19, machine learning on chest CTs has the potential to be helpful for: (a) triage in resource-constrained environments (e.g. diagnosis model), (b) evaluating complications, or (c) prediction of worsening vs. improvement.
- Any machine learning models developed for medical applications should only be deployed in the real world after extensive validation, in agreement with medical best practices, and under the guidance of medical professionals.
About the Featured Image
The CT slices in the featured image are from Radiology Assistant. The ROC curves and upper model figure are from Mei et al. The middle model figure is from Li et al.
Want to be the first to hear about my articles bridging healthcare, artificial intelligence, and business—and get a free list of my favorite health AI resources? Sign up here.