In this post, I discuss L1, L2, elastic net, and group lasso regularization on neural networks. I describe how regularization can help you build models that are more useful and interpretable, and I include Tensorflow code for each type of regularization. Finally, I provide a detailed case study demonstrating the effects of regularization on neural network models applied to real clinical and genetic data from the Framingham study.
Benefits of Regularization
Regularization Can Make Models More Useful by Reducing Overfitting
Regularization can improve your neural network’s performance on unseen data by reducing overfitting.
Overfitting is a phenomenon where a neural network starts to memorize unique quirks of the training data (e.g. training data noise) instead of learning generally-applicable principles. A model that has “overfit” will get high performance on the training data but bad performance on the held-out test data, meaning the model won’t be useful in the real world because it won’t perform well on data it has never seen before. Since the whole point of neural network models is to solve real problems on new data, we want to avoid overfitting so that we obtain a practically useful model.
Here is an example of overfitting in a classification problem:
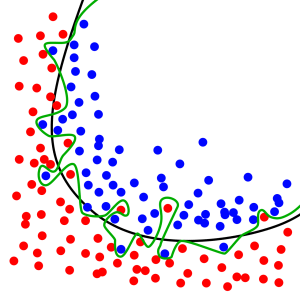
In this classification task we want to learn a line that separates the red dots from the blue dots. The black line represents a good classifier that appears to capture the general principle of where the blue dots and red dots are in the space. The green line represents a classifier that has suffered from overfitting because it’s too specific to this exact training set of red and blue dots.
Observe how the green classifier line does worse on an unseen test set, even though the unseen test set is following the same general layout of blue and red dots. Also notice that the black classifier line, which has not overfit, still works well on the test set:
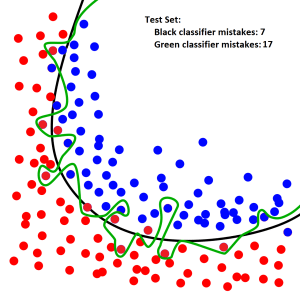
The most extreme version of overfitting happens when a neural network memorizes all of the examples in the training set. This can happen when the neural network has many parameters and is trained for too long.
Regularization Can Make Models More Understandable
Specifically, “lasso” regularization tries to force some of the weights in the model to be zero.
- In regression, one weight corresponds to one variable, so lasso regularization (using an L1 penalty) can directly “zeros out” a certain input variables by “zeroing out” the corresponding weight.
- In neural networks, we need to use “group lasso” regularization in order to zero out entire input variables and obtain a more interpretable model. This is because neural networks apply many weights to a single input variable, so we must consider all these weights as a “group.” (See later sections for more details.)
Silly example:
- We want to build a model to predict diabetes risk based on a clinical data set with these variables: [height, age, past diagnoses, past medications, favorite color, favorite animal]
- We build the model incorporating lasso regularization and discover that the model has chosen to “zero out” the weights corresponding to “favorite color” and “favorite animal,” which tells us that these variables are not useful in predicting future diagnosis of diabetes.
How to Regularize Neural Network Models
General Setup
Here’s the general setup of regularization for a neural network model, where W represents the weights of your neural network model:

Every neural network has a loss function that is used in training to adjust the neural network’s weights. The loss function measures how different the neural network’s predictions are from the truth (see this post for review.)
Regularization merely adds a “regularization term” (shown above in blue) to the loss function.
- λ (lambda) determines how strongly the regularization will influence the network’s training.
- If you set λ=0, there is no regularization at all, because you’ve zeroed out the entire regularization term.
- If you set λ= 1,000,000,000 then that would be extremely strong regularization, which would make it hard for your model to learn anything.
- Reasonable choices of the regularization strength λ could be 0.001, 0.01, 0.1, 1.0, etc. In the end, the best λ value to use depends on your data set and the problem you are solving. You can check the performance of your model with different λ strengths on your validation set, and then choose the λ that gives the best result. Thus, the regularization strength λ becomes a hyperparameter of your model that you can tune on the validation set.
- math
represents the actual regularization operation. We’re calling the weights of the neural network W, so regularization is just a mathematical operation on the weights. Different kinds of regularization correspond to different mathematical operations.
L1 Regularization
Here’s the formula for L1 regularization (first as hacky shorthand and then more precisely):
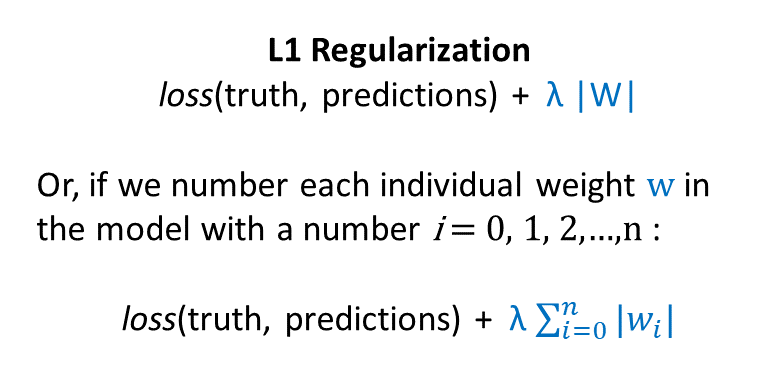

Thus, L1 regularization adds in a penalty for having weights of large absolute value. L1 regularization encourages your model to make as many weights zero as possible.
Here’s an example of how to calculate the L1 regularization penalty on a tiny neural network with only one layer, described by a 2 x 2 weight matrix:
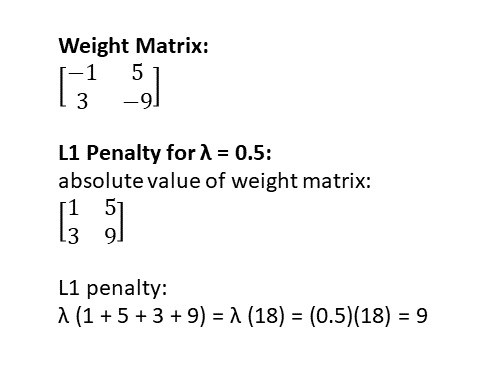

When applying L1 regularization to regression, it’s called “lasso regression.”
Here’s Tensorflow code for calculating the L1 regularization penalty for a weight matrix called weights:
l1_penalty = tf.reduce_sum(tf.abs(weights))
L2 Regularization
Here’s the formula for L2 regularization (first as hacky shorthand and then more precisely):

Thus, L2 regularization adds in a penalty for having many big weights. L2 regularization encourages the model to choose weights of small magnitude.
Here’s an example of how to calculate the L2 regularization penalty on a tiny neural network with only one layer, described by a 2 x 2 weight matrix:
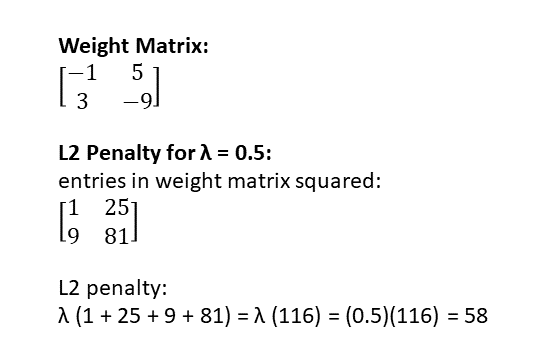

If you apply L2 regularization to regression, it’s referred to as “ridge regression.”
Here’s Tensorflow code for calculating the L2 regularization penalty for a weight matrix called weights:
l2_penalty = tf.reduce_sum(tf.nn.l2_loss(weights))
The Tensorflow function “l2_loss” calculates the squared L2 norm. The squared L2 norm is another way to write L2 regularization:
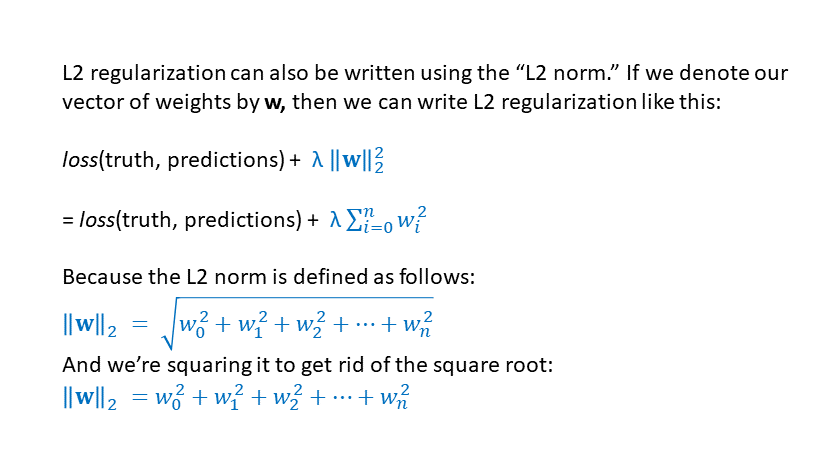

Comparison of L1 and L2 Regularization
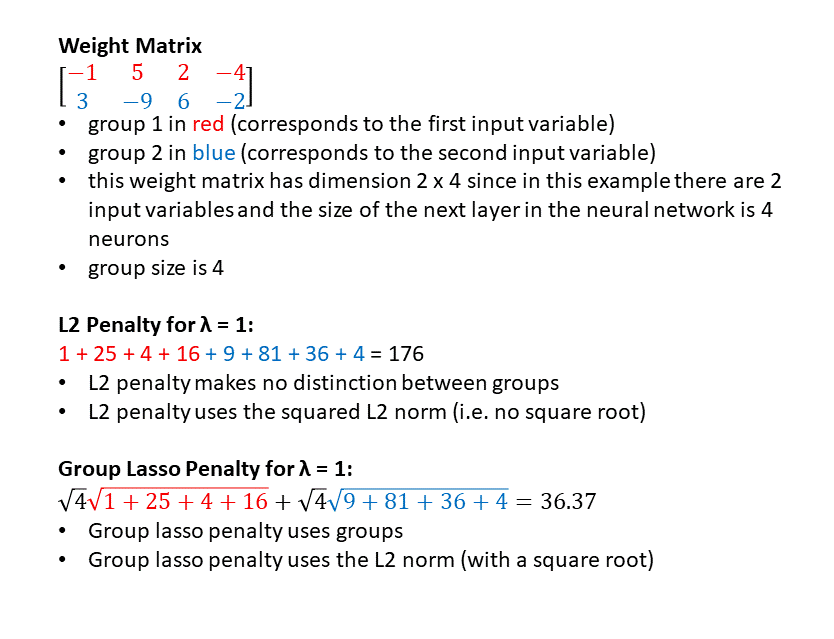
Notice that in L1 regularization a weight of -9 gets a penalty of 9 but in L2 regularization a weight of -9 gets a penalty of 81 — thus, bigger magnitude weights are punished much more severely in L2 regularization.
Also notice that in L1 regularization a weight of 0.5 gets a penalty of 0.5 but in L2 regularization a weight of 0.5 gets a penalty of (0.5)(0.5) = 0.25 — thus, in L1 regularization there is still a push to squish even small weights towards zero, more so than in L2 regularization.
This is why L1 regularization encourages the model to make as many weights zero as possible, while L2 regularization encourages the model to make all the weights as small as possible (but not necessarily zero).
Elastic Net Regularization
“Elastic net regularization” sounds fancy, but it simply means using both L1 and L2 regularization at the same time:
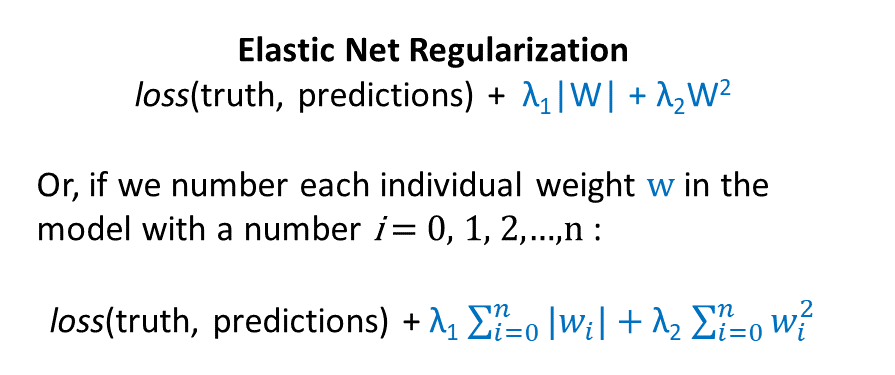

Here, we have two lambdas: one that controls the strength of the L1 regularization term, and another that controls the strength of the L2 regularization term. Both of these lambda values can be tuned using the validation set, as described previously.
Group Lasso Regularization
Group Lasso was introduced by Yuan and Lin in 2006:
(That paper has since been cited over 5,000 times.)
What is group lasso? Recall that L1 regularization is sometimes called “lasso regularization” and the purpose is to zero out some of the variables. In a similar vein, “group lasso” is a technique that allows you to zero out entire groups of variables. All members of a particular variable group are either included in the model together, or excluded from the model (zeroed out) together.
Here are two situations where group lasso is particularly useful:
For Categorical Variables: If you’ve represented a categorical variable as a one-hot vector — i.e., as a collection of binary covariates — group lasso can ensure that all of the binary covariates corresponding to a single categorical variable get “zeroed out” or “kept” together. For example, if you have a categorical variable “color” with possible values “red,” “blue,” and “green,” then you can represent this categorical variable with a one-hot vector of length three, corresponding to splitting a single “color” column into three binary columns: “red yes/no,” “blue yes/no,” and “green yes/no.” Group lasso can help you zero out all three columns together, or keep all three columns, treating them as a single unit.
For Neural Networks: If you’re training a neural network, group lasso can “zero out” entire input variables, and help you obtain a more interpretable model.
- As described previously, good old L1 regularization can zero out entire variables in regression easily, since it only has to zero out one weight to zero out one variable.
- But in a neural network, many weights act on one variable, which means we have to zero out all those weights at the same time in order to zero out that variable. Group lasso lets us group together all the weights corresponding to one variable, to achieve this goal.
The figure below shows “XW”: a 2-dimensional data input matrix X multiplied by a neural network weight matrix W. In this case W maps the 2-dimensional input to a 4-dimensional hidden layer. The input X consists of “Patient A” with variables “Blood Pressure” and “Cholesterol” (which have been appropriately normalized as described here.) You can see the first row of the weight matrix highlighted in red, which corresponds to weights that will multiply the blood pressure variable. The second row of the weight matrix is highlighted in blue, which corresponds to weights that will multiply the cholesterol variable. Thus, if we want to “zero out” the variable blood pressure, we need to “zero out” all four weights in the top row of the weight matrix.
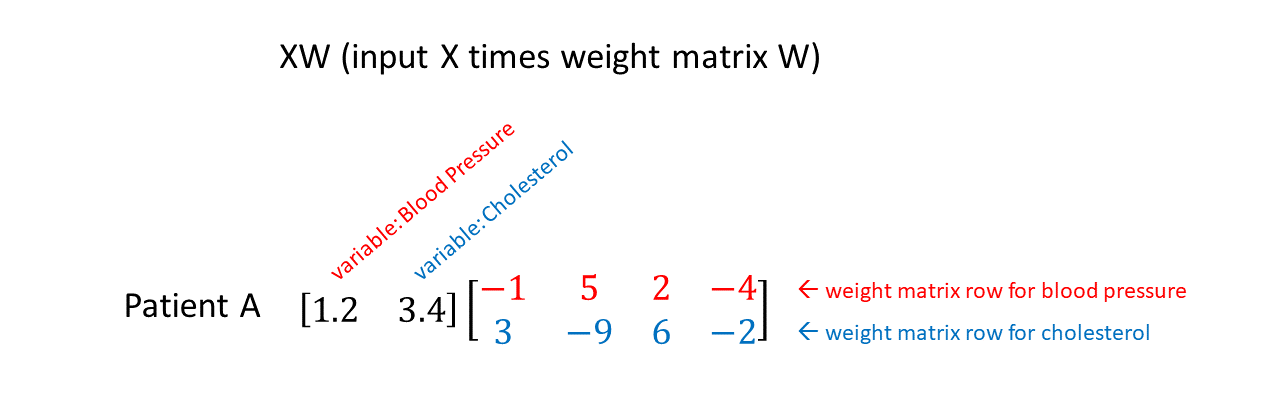
(Note: in most papers, neural network math is written as WX+b, where W is the weight matrix, X is the input, and b is the bias vector. Why am I writing XW here? Well…In Tensorflow, the implementation of a fully connected layer uses XW instead of WX. See this post for additional comments on notation in theory vs. practice.)
Here’s a paper describing group lasso applied to neural networks:
and here’s a Bitbucket repository for the paper, where the authors explain,
For each node in the network, we include a regularization term pushing the entire row of outgoing weights to be zero simultaneously. This is done by constraining the L2 norm of the row, weighted by the square root of its dimensionality.
Here’s the formula for group lasso regularization:
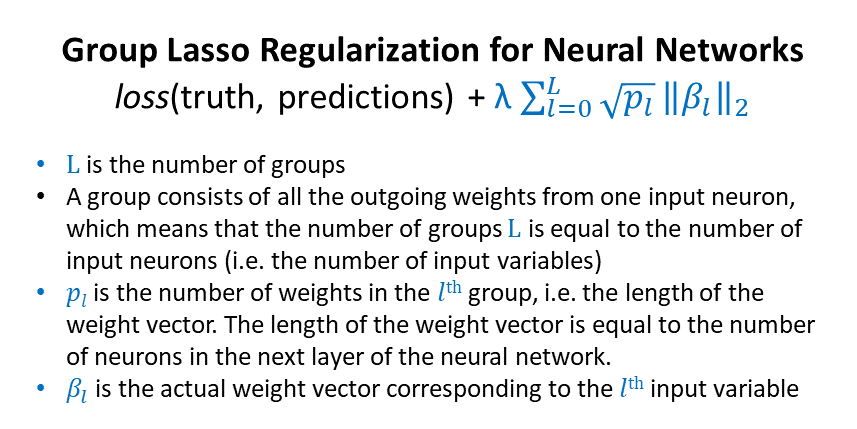

Here’s one Tensorflow function for the group lasso penalty, from this repository:
# Define the group lasso penaltydef groupl1(x):returnT.sum(T.sqrt(x.shape[1])*T.sqrt(T.sum(x**2, axis=1)))
Here’s another Tensorflow implementation of the group lasso penalty:
euclidean_norm =tf.sqrt( tf.reduce_sum(tf.square(weights),axis =1) )
#Must cast num_outputs to a float in order for tensorflow to take the square root
account_for_group_size = tf.sqrt( tf.cast(num_outputs, dtype=tf.float32) )
penalty = tf.reduce_sum( tf.multiply(account_for_group_size, euclidean_norm) )
Somewhat confusingly, group lasso in this paper uses the L2 norm like in L2 regularization (summing up the squared values of the elements). But traditional “lasso” uses the L1 norm! What’s going on here?
Wikipedia defines “Lasso” as:
Lasso (least absolute shrinkage and selection operator) is a regression analysis method that performs both variable selection and regularization in order to enhance the prediction accuracy and interpretability of the statistical model it produces.
“Group lasso” is performing both variable selection (by zeroing out groups of weights corresponding to particular input variables) and regularization. Also, even though group lasso includes an L2 norm, it is NOT the same as L2 regularization:

Framingham Case Study
Background
The Framingham Heart Study started in 1948 and has continued to this day. The Framingham study is responsible for much modern knowledge about heart disease, including that eating a healthy diet, maintaining a healthy weight, not smoking, and engaging in regular exercise can reduce the risk of heart disease. The Framingham heart study data set includes clinical variables (e.g. age, smoking status), genetic variables, and heart disease outcomes (e.g. whether or not a patient had a heart attack.)
In summer 2018, I spent a few months analyzing part of the Framingham heart study data to determine whether predictive models that included both clinical and genetic data would perform better than predictive models built using clinical data alone. I suspected that it would be difficult to see any benefit in predictive performance by adding genetic data on top of clinical data, because:
- Many clinical variables (e.g. cholesterol levels, triglyceride levels, blood pressure) summarize both lifestyle choices (e.g. diet, exercise, substance use) and genetics. Therefore, clinical data includes already genetics, albeit implicitly.
- There are thousands of spots in the human genome that influence heart disease. These spots are spread out across the entire genome. Each spot contributes a only a small amount to heart disease risk on its own. However, about 50% of the risk of heart disease is genetic — which means that in aggregate, all these spots together have a noticeable effect on heart disease risk. Because there are so many different contributing spots, each with small effect, we need a HUGE sample size to pick up on patterns useful for making predictions. Unfortunately, in my subset of the Framingham data, I had 500,000 genetic variables but only 3,000 patients, which means the problem is underdetermined (more variables than training examples.)
Model Results
I trained various feedforward neural networks on a combination of clinical and genetic data from the Framingham data set to predict heart disease risk. I applied different kinds of regularization to the first layer weight matrix. The table below shows the performance of the different models (Acc = accuracy, AUC = area under the receiver operating characteristic, AP = average precision):
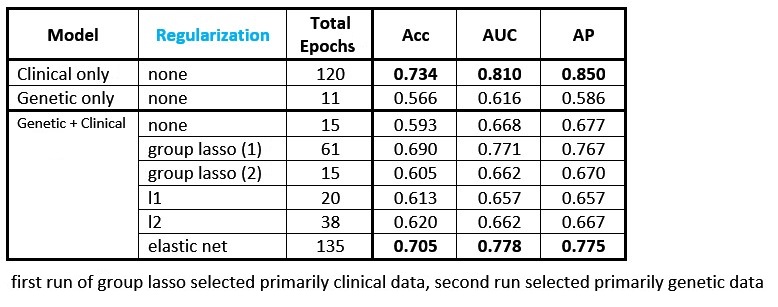
Key points about the results:
- The best-performing model uses clinical data only (first line.)
- The model using genetic data only (second line) has much lower performance than the model that uses clinical data alone.
- The models combining genetic and clinical data all get lower performance than the model using clinical data alone, likely because the sample size of 3,000 patients is not large enough to learn anything meaningful from the genetic data. The model instead overfits to noise in the genetic training data, and is then incapable of generalizing to the genetic data in the test set. This has the effect of “muddying” the test set performance (relative to the model where clinical variables are the only inputs.)
First-Layer Weight Matrix Heat Maps
We can gain even more insights by inspecting heat maps of the first-layer weight matrices for the different regularization approaches. Each row of the heat map corresponds to a different input variable. The top 20 rows (genetic_0 to genetic_19) correspond to weights applied to a learned 20-dimensional representation of the 500,000 genetic input variables. The bottom rows (from SYSBP1 to DIAB_1.0) correspond to weights applied to clinical variables.
Here are heat maps for the model with (A) no regularization, (B) L1 regularization, (C) L2 regularization, and (D) elastic net regularization:
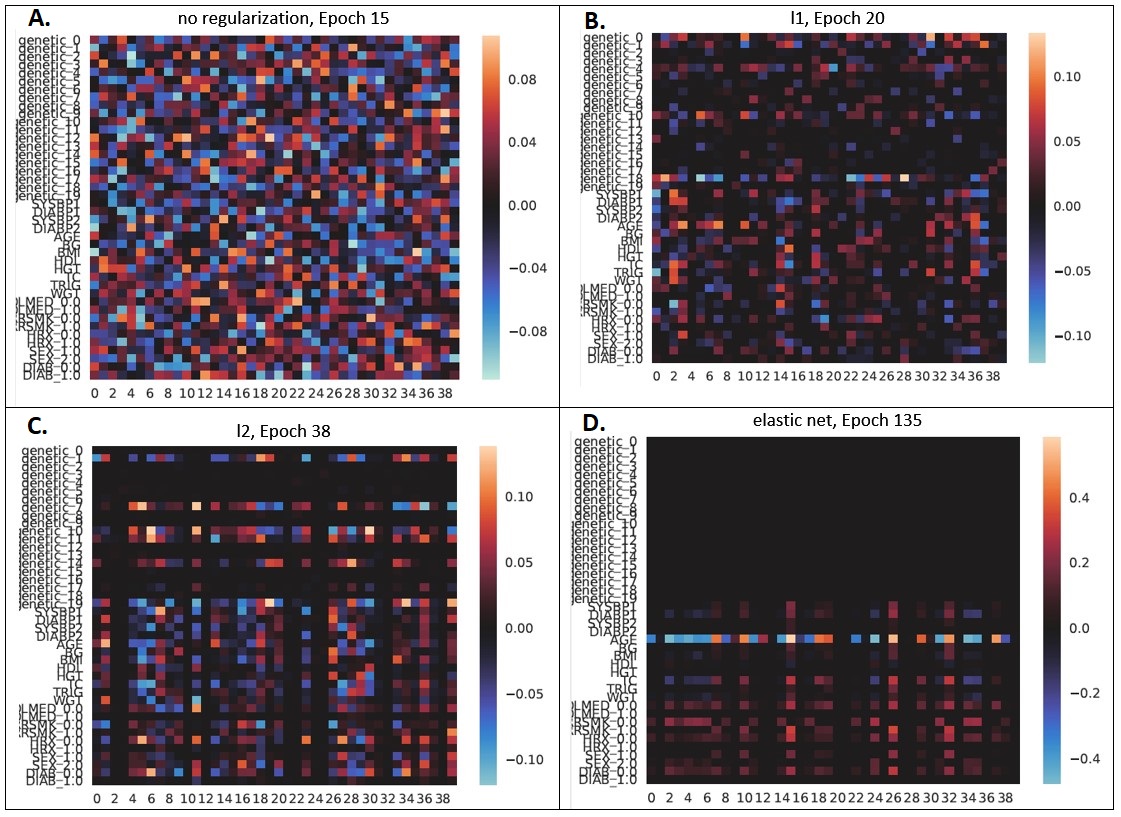
Panel A shows the first-layer weight matrix with no regularization applied. We can see that there are large-magnitude positive and negative weights scattered throughout with no particular pattern.
Panel B shows the first-layer weight matrix with L1 regularization applied. We can see that there are more zero-valued (black) weights. However, there is no particularly strong pattern at the level of variables (rows), which is to be expected as L1 regularization considers the absolute value of each weight independently.
Panel C shows the first-layer weight matrix with L2 regularization applied. This has a greater number of smaller magnitude weights than the unregularized case.
Panel D shows the first-layer weight matrix with elastic net regularization applied. For this data set, elastic net regularization ended up achieving the best performance of all the regularization methods considered. It also produced a pretty first-layer weight matrix: all the genetic variables are “zeroed out” and the brightest clinical variable is “age” (which is somewhat funny, since I was looking at ~40-year risk of heart disease, so if you start out at age 80 at the beginning of the study period you’re going to have higher risk than someone starting out at age 30.)
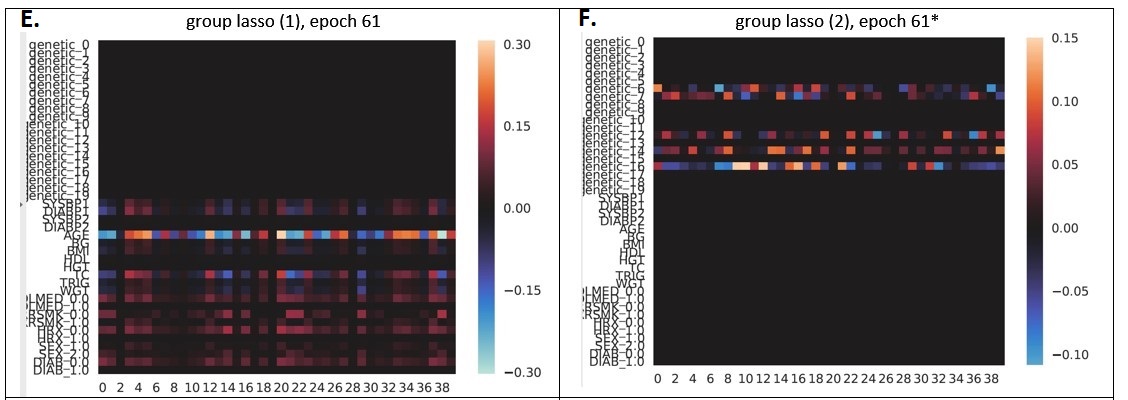
Finally, in Panels E and F we can see the results of two different models trained using group lasso regularization. The models were trained with different random initializations and ended up finding different solutions:
- The model in Panel E “zeroed out” all of the genetic variables, and kept most of the clinical variables (again, similar to the elastic net case, the highest-weighted clinical variable is “age”.) In the results table above this model is listed as “group lasso (1)” and achieves 0.690 accuracy, 0.771 AUROC, and 0.767 AP.
- The model in Panel F ultimately “zeroed out” all of the clinical variables, and attempted to use the genetic variables for prediction. The best validation performance was actually achieved at epoch 15, rather than epoch 61, because at epoch 15 the model hadn’t finished killing all the clinical variables yet. Performance at epoch 15 was 0.605 accuracy, 0.662 AUROC, and 0.670 AP, and it just got worse after the model decided it wanted to kill off the clinical variables.
Thus, this experiment also ended up being a nice demonstration of how neural networks with different random initializations can find different solutions on the same data set.
Note that in many cases, application of regularization leads to better test set performance, which is not explicitly illustrated in this example. If I had added regularization to the best-performing “clinical data only” model, perhaps it would have resulted in even higher performance.
Conclusions
- L1, L2, elastic net, and group lasso regularization can help improve a model’s performance on unseen data by reducing overfitting.
- L1 regularization in regression and group lasso regularization for neural networks can produce more understandable models by “zeroing out” certain input variables.
- Visualizing weight matrices with heat maps in regularized neural network models can provide insights into the effects of different regularization methods.
About the Featured Image
The featured image is an oil painting called “The Herd Quitter” by C.M. Russell of cowboys attempting to lasso a bull. A lasso for livestock is also called a “lariat,” “riata,” “reata,” or simply a “rope.”
Want to be the first to hear about my articles bridging healthcare, artificial intelligence, and business—and get a free list of my favorite health AI resources? Sign up here.


Comments are closed.