Your genome is approximately 750 megabytes of information (3 x 10^9 letters x 1 byte/4 letters). That’s about half the size of an operating system except it codes for an entire human body, and the entire code fits into a volume a hundred times smaller than a grain of rice. Your brain, which develops as specified by your genome, is an incredible supercomputer that also happens to require less power than a dim light bulb — literally tens of thousands of times more energy-efficient than manmade supercomputers.
So how does the genome work?
Genomics, transcriptomics, and proteomics are data-driven fields that aim to answer the question of how genomes code for a living, breathing person (or cat, or plant, or bacterium). This post introduces genomics, transcriptomics, and proteomics to computational scientists and engineers who are interested in working on “omics” data and would like a quick introduction to where this data comes from and the key biology behind the data. This post uses minimal jargon and multiple analogies to clarify the similarities and differences between these different kinds of “omics” data and how they provide different windows into one of the most complicated pieces of software in existence – your DNA.
The Cell
The cell is the basic building block of all living creatures: people, dogs, mice, trees, flowers, flies, and bacteria. All forms of “omics” are measuring large quantities of something found inside of cells.
Here is a picture of some cells taken with a microscope. I’ve outlined a few of the cells in red. Each red circle corresponds to one cell:
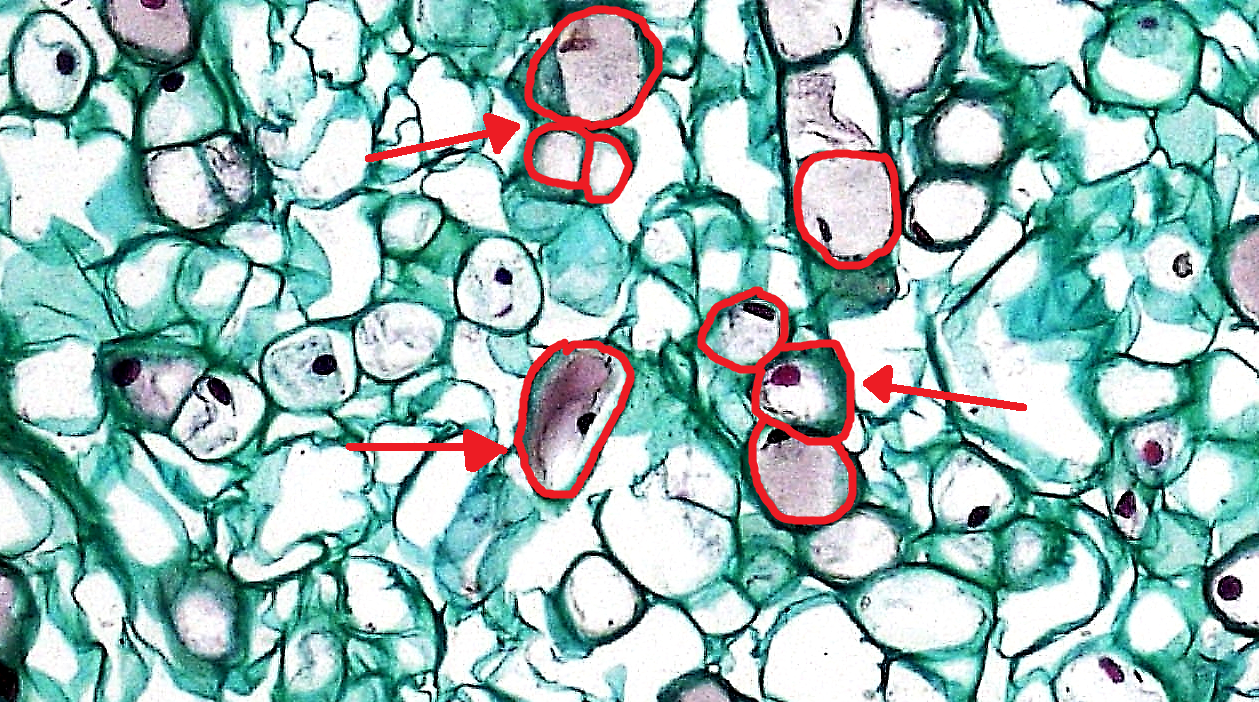
Modified from Wikipedia.
Robert Hooke discovered and named cells in 1665. He called them cells because under the microscope they looked like “cellula” or small rooms – in other words, “cells” are called “cells” because they can look like prison cells under a microscope.
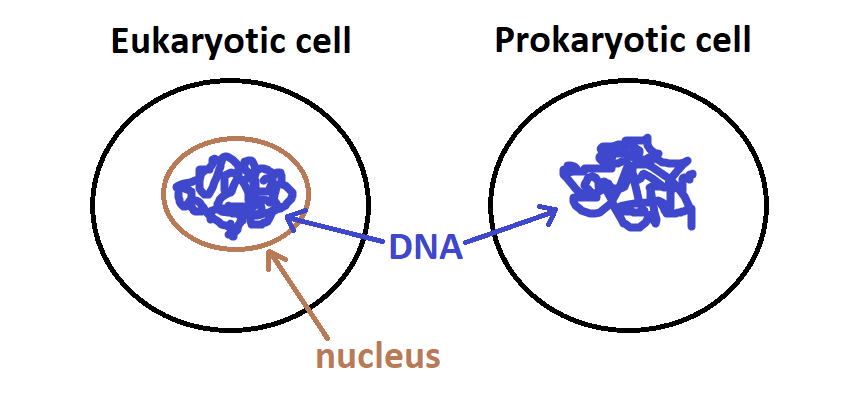
Cells from people, dogs, mice, trees, flowers, and flies are “eukaryotic” which means that these cells pack up their DNA in a bag (the nucleus).
Cells from bacteria are called “prokaryotic” because they do not keep their DNA in a bag (i.e. they have no nucleus):
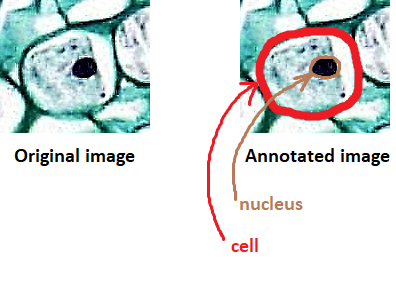
Here’s a close up of the earlier (eukaryotic) cell picture with the nucleus circled in brown:
DNA and Genes
Your DNA is a long sequence of 3 billion letters, found inside of your nucleus. Every cell in your body (with rare exception) has a full copy of your entire genome (i.e. all your DNA). Because there are about 37.2 trillion cells in your body, that means you are carrying around 37.2 trillion copies of your whole DNA sequence! The implications are interesting: this means that a cell in your fingertip contains all the instructions needed to build your entire brain.
A “gene” is a particular segment of DNA that encodes instructions for making a protein.
We can think of the body as a factory, which needs certain “machines” (proteins) to function:
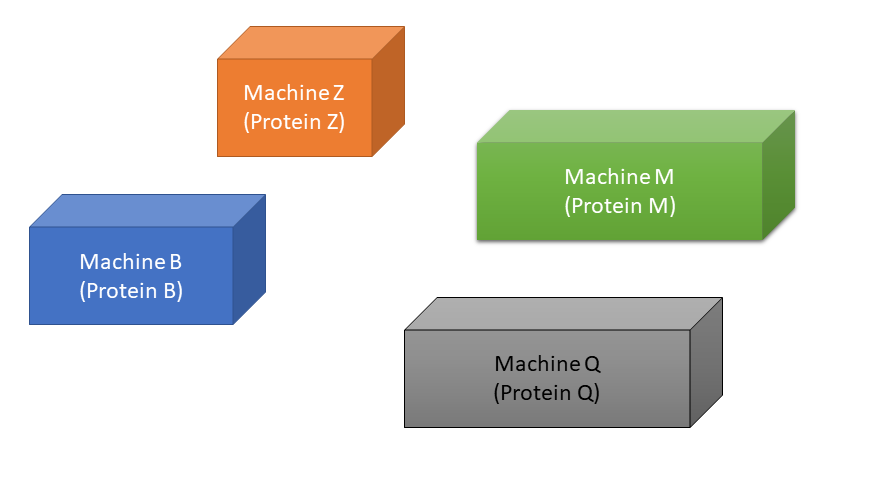
A gene is a set of instructions for how to build a particular machine. For example, Gene B contains instructions for building Machine B (Protein B), and Gene Q contains instructions for building Machine Q (Protein Q), etc.
Only 1% of the genome actually consists of genes. The genes are referred to as the “coding” part of the genome, since the genes “code” for proteins. There are approximately 25,000 genes in the human genome, corresponding to 25,000 little machines (proteins) that make your body function.
The remaining 99% of the genome is “noncoding.” The “noncoding” part of the genome is still important because it tells the cell how to make use of the “coding” part.
In other words, if the “coding” part serves as instructions for how to build different machines, the “noncoding” part serves as instructions for how to make use of those machines in the factory: when the machines need to be turned on, when the machines need to be turned off, how many machines are needed, when the machines should be used, etc.
If we picture the genome as a book, it will contain some text on how to build the machines (the coding part, shown in orange and blue below), and some text on how to use the machines (the noncoding part, shown in black below):
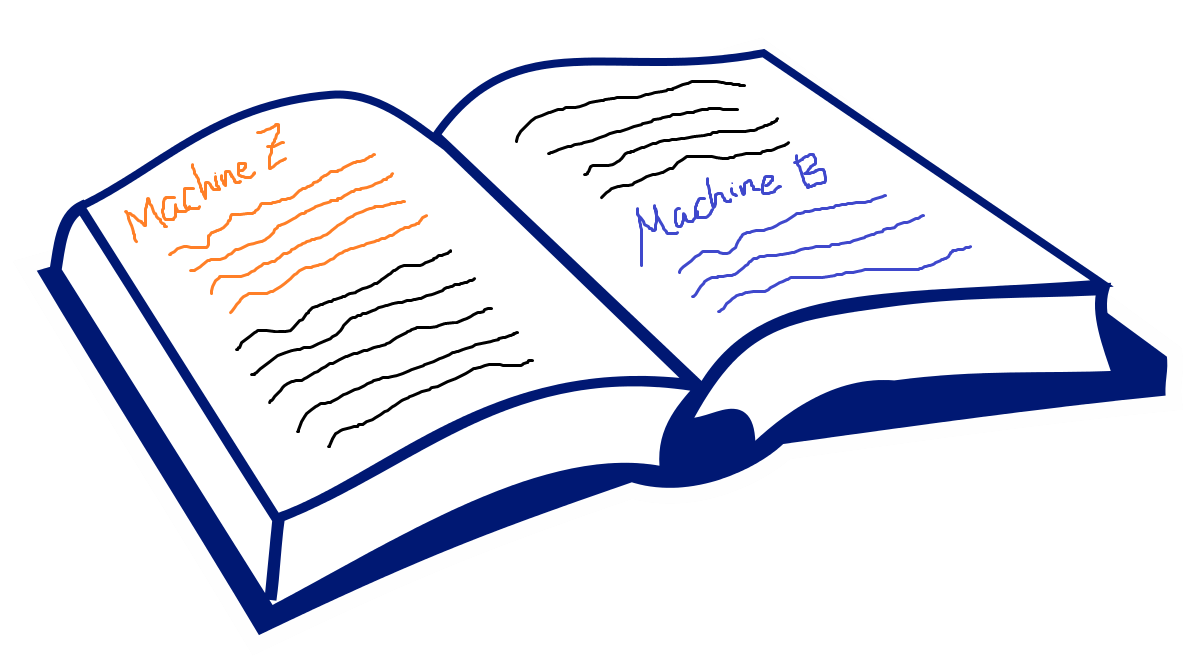

Modified from blank open book picture on Wikipedia
Nucleotides: The Building Blocks of DNA (A, T, C, G)
The 3 billion letters in DNA come from a limited alphabet of only 4 possible letters: A, T, C, and G. (Fun fact: this is why the movie “Gattaca” about the triumph of a genetically “inferior” man over his society uses only the letters A, T, C, and G in its title.)
Here is the entire DNA sequence for the human gene S100A10 (also called “p11”), one of the shortest genes in the genome:
CCGGTTACCTCTGGTTCTGCGCCACGTGCCCCACCGGCAGGACGGCCGGGTTCTTTGATT TGTACACTTTCTAAAACCAAACCCGAGAGGAAGGGCAGGCTCAGGGTGGGATGCCCTAAA TATTCGAGAGCAGGACCGTTTCTACTGAAGAGAAGTTTACAAGAACGCTCTGTCTGGGGC GGGCGAGCGCTCTGCGAGGCGGGTCCGGGAGCGAGGGCAGGGCGTGGGCCGCGCGCCCGG GGTCGGGGGAGTCGGGGGCAGGAAGAGGGGGAGGAGACAGGGCTGGGGGAGCGCCCTGCC GAGCGCCCGCCAGGCTCCTCCCCGTCCCGCACCGCCTCCCTCTACCCACCCGCCGCACGT ACTAAGGAAGGCGCACAGCCCGCCGCGCTCGCCTCTCCGCCCCGCGTCCACGTCGCCCAC GTCGCCCAGCTCGCCCAGCGTCCGCCGCGCCTCGGCCAAGGTGAGCTCCCAGTTCGGCCC
Sequence source: ebi.ac.uk
The building blocks of DNA (the four-letter alphabet A, T, C, and G) are called “nucleotides.” Their full names are guanine (G), adenine (A), cytosine (C) and thymine (T).
Each nucleotide building block is in turn made from carbon, hydrogen, oxygen, and nitrogen. For example, cytosine is made of 13 atoms. Here is a 2D “stick and ball cartoon” of what cytosine looks like:

Based off of chemical structure shown at cs.boisestate.edu
And here is a 3D rendering of what cytosine looks like:
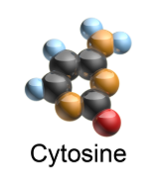
3D rendering source: Wikipedia
(Technical note: the pictures of cytosine above are actually of the “nucleobase” cytosine which is the most fundamental chemical part. A “nucleoside” is a nucleobase plus the sugar ribose. A “nucleotide” – the direct building block of DNA – is a nucleobase plus the sugar ribose plus one or more phosphate groups. So, technically, in order for the picture of cytosine shown above to represent a true building block of DNA, it needs to have a ribose and a phosphate added.)
When we string together the nucleotides A, T, C, and G to make DNA, we get something called a “nucleic acid” – hence the full name for DNA, “deoxyribonucleic acid.“
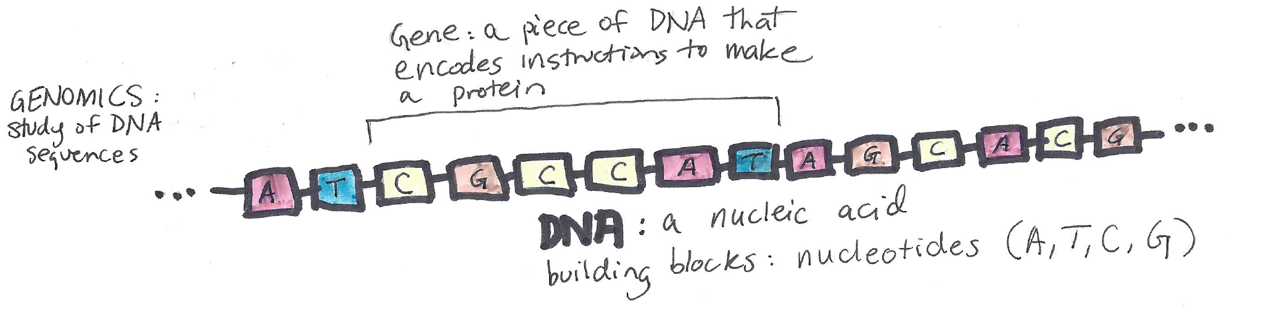
DNA is double stranded; for simplicity, I only showed one strand in the cartoon graphic above. The double-stranded nature of DNA is the reason for the “double helix”:
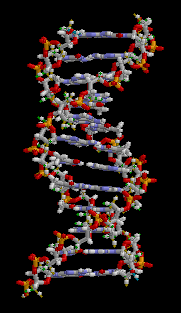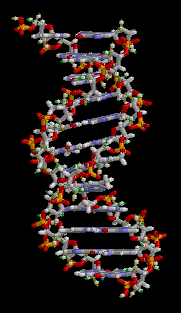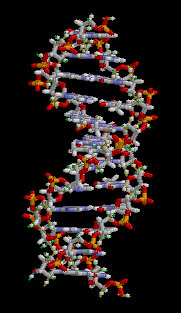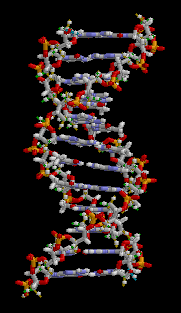
Animation from Wikipedia
The two strands of DNA wind around each other to form a double helix. The information in one strand is a “mirror copy” of the information in the other strand. This is due to how the A, T, C, and G building blocks pair with each other. A pairs with T, and C pairs with G. Therefore, if one strand reads “AAATTC” the opposite strand will always read “TTTAAG”. The information content of one strand is equal to the information content of another strand.
It might seem like a waste to have two copies of the same information, but in fact the double-stranded nature of DNA is important:
- Double-strandedness facilitates DNA replication (i.e. copying of the entire genome, which must take place every time one cell divides into two cells, as each “daughter cell” needs a full copy of the genome) ;
- Double-strandedness enables the body to error-correct the genome (i.e. if there is a mistake made in the copying process, double-strandedness enables correction of the errors);
- Double strands are more stable than single strands and it’s very important for the blueprint of life to be stable.
From a computational perspective, you only need the sequence of one strand to perform analyses.
The Human Genome Project was an international project to figure out the complete DNA sequence of a human being – i.e., read out all 3 billion A, T, C, and G letters in one person’s genome. The Human Genome Project cost $2.7 billion dollars. Nowadays, it is possible to get a complete genome sequence for orders of magnitude less money – for example, you can get your entire genome sequenced by Nebula Genomics for $299.
Genomics
Genomics is the study of DNA sequences – in humans, in animals, in bacteria, etc. Often genomes are compared to one another in order to learn about health and disease.
For example:
- Genomes of healthy and sick people can be compared to learn about genes that influence disease risk or directly cause disease (e.g. see “genome-wide association study“).
- Genomes of many people can be compared to learn about natural variation in the genome unrelated to disease – e.g. genes that influence hair color, eye color, height, and other characteristics.
- Genomes of people and animals can be compared in order to reconstruct the tree of life and understand evolutionary history.
Genetic Differences and Similarities
Any randomly-selected man on the planet is 99.9% genetically identical to any randomly-selected woman on the planet.

For example, George Takei and Oprah Winfrey are 99.9% genetically identical
Chimpanzees and humans are 96% genetically identical overall (and 98% identical in the coding regions):

Dogs and wolves are 99.9% identical and in fact are so genetically similar that they are technically the same species:

Wolf image from Wikipedia; chihuahua image from Wikipedia.
Thus, much of genomics is about identifying tiny portions of the genome that lead to differences (with dogs being a particularly interesting species to study, since they vary so much in size, color, behavior, longevity, and disease risk). Genomics can be “searching for a needle in a haystack” especially when comparing different human genomes to each other in order to understand disease risks, as any given pair of human beings is mostly identical.
Kinds of Genomic Data
Hypothetically the DNA sequence for one human could be stored in about 750 megabytes. However, in reality the process of DNA sequencing isn’t perfect, and instead of getting one perfect readout of the entire genome start to finish, it instead produces many partial reads which must be assembled together like a puzzle. For example let’s say you were given the following sentence fragments:
- “the bird flies in”
- “flies in the sky”
- “in the sky and sings”
By aligning these sentences and overlaying them, you can reconstruct the sentence “The bird flies in the sky and sings.”
Because DNA is sequenced in this manner, a raw DNA sequence straight off the sequencing machine is about 200 gigabytes, and must be “cleaned up” before analysis. For more information you can check out this post.
So far, we have only been talking about getting the complete genome sequence, something that is referred to as “whole genome sequencing.” It’s also possible to do genomic studies with other kinds of genomic measurements:
- Whole exome sequencing (“WES”): in this technique, only the coding parts of the genome (the parts that code for proteins) are sequenced.
- SNP genotyping: in this technique, only single letters at locations that are known to be important are measured. A “SNP” is a “single nucleotide polymorphism”: single because it’s only one location (e.g. “position 4,576,877”), nucleotide because the building blocks of DNA are nucleotides and there is one nucleotide at that location (A, T, C, or G), and polymorphism because these are special locations being considered which are known to vary in the population (i.e. they are “polymorphic”). Having certain nucleotides at a SNP rather than others can result in dramatic consequences. For example, the disease sickle cell anemia can be caused by a single nucleotide change.
RNA & Transcription
Like DNA, RNA is a nucleic acid – specifically, ribonucleic acid.
Comparison of DNA and RNA:
- DNA is made from A, T, C, and G. RNA is made from A, U, C, and G. Thus, RNA contains U (uracil) instead of T (thymine).
- DNA is double-stranded. RNA is single-stranded.
- DNA is for stable, long-term storage of all the information needed to make a living creature. RNA is used as a temporary template to aid in the process of making proteins.
Here’s the RNA sequence that can be made off of the the human gene S100A10:
AAUCAAAGAACCCGGCCGUCCUGCCGGUGGGGCACGUGGC GCAGAACCAGAGGUAACCGGUUUAGGGCAUCCCACCCUGA GCCUGCCCUUCCUCUCGGGUUUGGUUUUAGAAAGUGUACA GCCCCAGACAGAGCGUUCUUGUAAACUUCUCUUCAGUAGA AACGGUCCUGCUCUCGAAUACCGGGCGCGCGGCCCACGCC CUGCCCUCGCUCCCGGACCCGCCUCGCAGAGCGCUCGCCC GGCAGGGCGCUCCCCCAGCCCUGUCUCCUCCCCCUCUUCC UGCCCCCGACUCCCCCGACCACGUGCGGCGGGUGGGUAGA GGGAGGCGGUGCGGGACGGGGAGGAGCCUGGCGGGCGCUC GUGGGCGACGUGGACGCGGGGCGGAGAGGCGAGCGCGGCG GGCUGUGCGCCUUCCUUAGUGGGCCGAACUGGGAGCUCAC CUUGGCCGAGGCGCGGCGGACGCUGGGCGAGCUGGGCGAC
Calculated using arep.med.harvard.edu
RNA is used as a template to build proteins. First, a piece of RNA is built based off of a gene in DNA. The process of making RNA off of DNA is called “transcription” because the RNA is “transcribed” off of the DNA, making use of the pairing rules discussed earlier (except this time instead of being a DNA strand-DNA strand pairing to form a double helix, it’s a DNA strand – RNA strand pairing to make an RNA template):
- RNA C pairs with DNA G
- RNA G pairs with DNA C
- RNA U pairs with DNA A
- RNA A pairs with DNA T
Next, that piece of RNA is used to make a protein, in a process called “translation” because the nucleotide building blocks of RNA are “translated” into the amino acid building blocks of proteins:
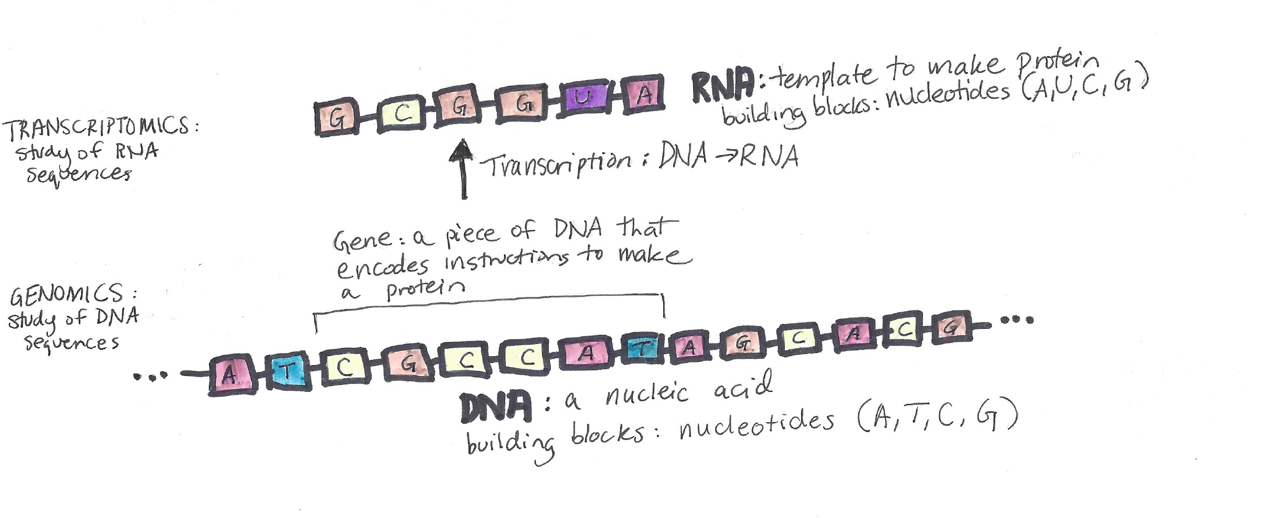
Transcriptomics
“Transcriptomics” is the study of RNA sequences. Transcriptomics is interesting because it tells us what genes are “turned on” in different parts of the body.
Let’s go back to the analogy of the DNA sequence as an instruction manual:

Recall that the orange and blue text contains instructions to build “Machine Z” and “Machine B” respectively (representing two genes, which provide instructions to build Protein Z and Protein B).
Now let’s say that we have to photocopy the page for “Machine Z” every time we want to add a new Machine Z to a room of our factory — maybe we’re handing off the photocopied page of Machine Z instructions to some worker who is going to go build Machine Z. If we photocopy the page for “Machine Z” a total of 25 times, then someone who sneaks in to the room of the factory and counts up the total number of photocopies for the Machine Z page can infer that in that particular room of the factory we need 25 Machine Z’s.
Thus, by counting up the number of photocopies (RNA transcripts) in a certain room of the factory (body part), we can infer which machines (proteins) are used the most in that part of the factory.
The body is made up of tissues – for example, muscle is a kind of tissue, and nerve cells are another kind of tissue. Here is a diagram from genevisible showing the top 10 tissues in which the gene S100A10 is “expressed” – in other words the tissues in which the gene S100A10 is transcribed most frequently into RNA:

We can see from the diagram that the gene S100A10 is used more in certain parts of the body than others. “Expression” is just another word for making a piece of RNA off of DNA. We say that a gene is “expressed” when that gene is used to make RNA templates. The phrase “measuring gene expression” refers to detecting the degree to which RNA is made from particular genes of interest. In other words, “measuring gene expression” is the same as “measuring the transcriptome.”
Proteins & Translation
So far, we have discussed how DNA gets made into RNA. Now we will discuss what RNA is used for: serving as a template to make proteins.
Proteins are little machines that carry out different jobs in the body. I do not use the word “machines” lightly – protein really are molecular machines. Here is an animation showing the motor protein kinesin dragging a vesicle (a kind of microscopic storage container, like a backpack) along a microtuble (a kind of microscopic support structure, like a wooden beam in a building):
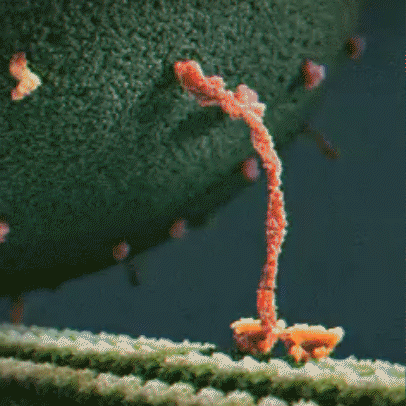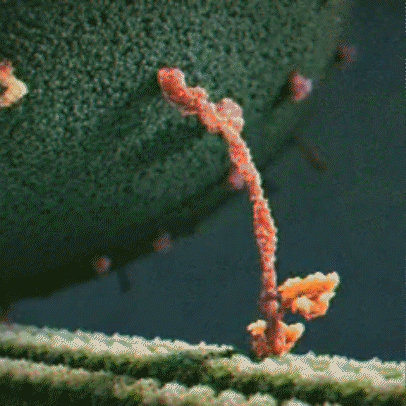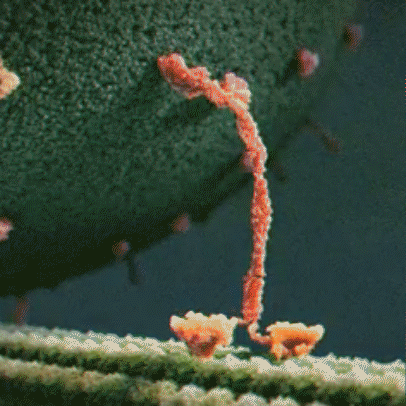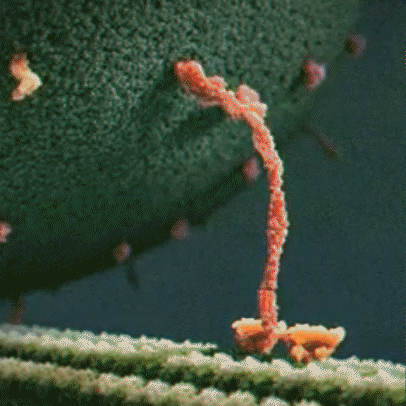
Proteins are made out of building blocks called amino acids. In humans, there are 20 amino acids available as building blocks for proteins.
Here are the 20 amino acids used to make proteins in humans, along with their single-letter and triple-letter abbreviations:
- G – Glycine (Gly)
- P – Proline (Pro)
- A – Alanine (Ala)
- V – Valine (Val)
- L – Leucine (Leu)
- I – Isoleucine (Ile)
- M – Methionine (Met)
- C – Cysteine (Cys)
- F – Phenylalanine (Phe)
- Y – Tyrosine (Tyr)
- W – Tryptophan (Trp)
- H – Histidine (His)
- K – Lysine (Lys)
- R – Arginine (Arg)
- Q – Glutamine (Gln)
- N – Asparagine (Asn)
- E – Glutamic Acid (Glu)
- D – Aspartic Acid (Asp)
- S – Serine (Ser)
- T – Threonine (Thr)
Going back to our S100A10 example, here is the entire protein sequence for the protein ultimately produced from the gene S100A10:
MPSQMEHAMETMMFTFHKFAGDKGYLTKEDLRVLMEKEFPGFLENQKDPLAVDKIMKDLD QCRDGKVGFQSFFSLIAGLTIACNDYFVVHMKQKGKK
Sequence source: uniprot.org
Reminder:
- When you read off a protein sequence, “G” stands for the amino acid “Glycine” but when you read off a DNA sequence, “G” stands for the nucleotide “Guanine.”
- Similarly, “C” represents “Cysteine” in a protein sequence but “Cytosine” in a DNA sequence,
- “T” represents “Threonine” in a protein sequence but “Thymine” in a DNA sequence,
- and “A” represents “Alanine” in a protein sequence but “Adenine” in a DNA sequence.
- Luckily, you can tell pretty quickly whether a sequence is for protein or DNA because the protein sequence will include way more letters than just ATCG.
In spite of the confusingly similar-sounding names, amino acids and nucleotides are very different from one another, with different molecular structures. Here are diagrams of the 20 amino acids (where O is oxygen, H is hydrogen, N is nitrogen, and angles without a letter are carbon):
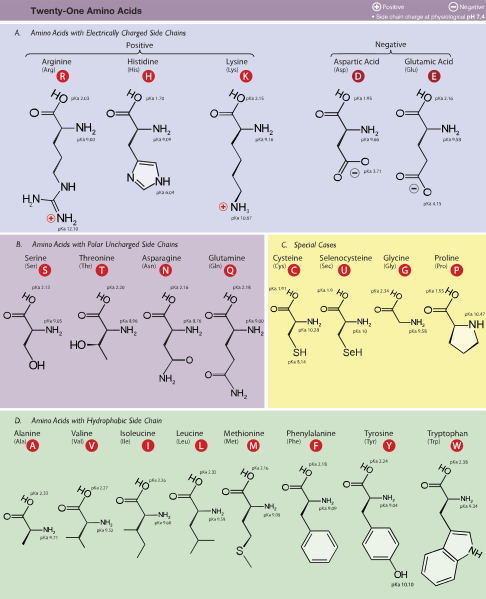
Source: Wikipedia
A protein is made using a piece of RNA as a template. The RNA template specifies what amino acids should be used to make the protein.
You might ask, how is it possible to specify instructions for a protein made of 20 different amino acids if RNA is made of only 4 kinds of nucleotides? If there were a one-to-one mapping between nucleotides and amino acids, then we’d only be able to code for 4 different amino acids. If there were a two-to-one mapping between nucleotides and amino acids, then we’d only be able to code for 4 x 4 = 16 different amino acids using these 2-element nucleotide codes:

Because two still isn’t enough, the body uses 3 nucleotides to indicate one amino acid. Using 3 amino acids provides 4 x 4 x 4 = 64 possibilities, which is plenty to code for the 20 available amino acids (with some triplets left over to code for “start” and “stop”). The grouping of 3 amino acids is referred to as a “triplet” or as a “codon.” During the process of “translation” (making a protein from RNA), the body reads the RNA in chunks of three letters, to produce one amino acid at a time.
The “breaking of the genetic code” occurred in 1966, and refers to the culmination of a painstaking effort to experimentally determine which amino acids corresponded to which nucleotide triplets.
Here is the genetic code, showing the mapping from RNA triplets (e.g. “UUU”) to amino acids (e.g. “Phe” which is “Phenylalanine”):

The sequence “AUG” (labeled in red) serves both as a “Start” indicator (“start codon”) and as the code for the amino acid methionine (Met). The sequences “UAA”, “UAG”, and “UGA” all code for “Stop.” The remaining three-letter codes all indicate a particular amino acid. You can see from the diagram that the genetic code is redundant, i.e. there are multiple ways to code for the same amino acid.
Here is our diagram with all the pieces in it: DNA, RNA, and protein.
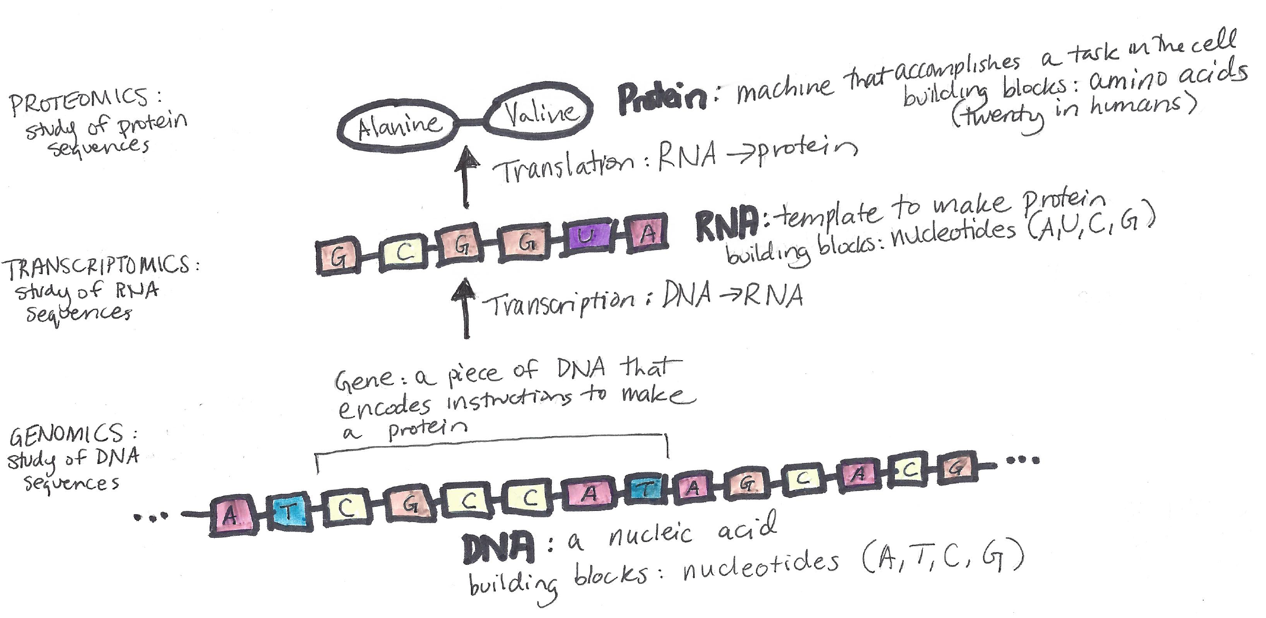
Notice that the DNA sequence labeled as a “gene” has the sequence CGCCAT, which according to the pairing rules described earlier, corresponds to the RNA sequence shown: GCGGUA. Then, using the genetic code table, we can see that GCG in the RNA corresponds to the amino acid alanine (Ala), while the GUA in the RNA corresponds to the amino acid valine (Val). (For simplicity, I left off “start” and “stop” codons in the diagram).
A quick side note on actual sizes: real genes are much bigger than the 6 nucleotides shown in this picture. The average protein-coding gene consists of 53,600 nucleotides (also written as “53.6 kb” where kb=kilobases; 53.6 x 1,000 bases). The largest gene is 2.4 megabases (dystrophin), which as you can imagine codes for a truly massive piece of RNA and ultimately a truly massive protein.
Visualizing Proteins
There are many ways to visualize the protein that is ultimately produced from a given gene. Proteins are produced as a sequence of amino acids, but they ultimately fold up into a 3D structure that is important for their functionality. It is a lot of work to determine the 3D structure of a protein, and a lot of work to determine a protein’s function. X-ray crystallography is one technique that can be used to determine the 3D structure of a protein. There is also a large area of computational research focused on predicting the 3D structure of a protein given its sequence alone (this is a very difficult problem).
Here are several visualizations of the protein produced from the S100A10 gene, from Swiss-Model. If you want to move around these renderings in an interactive 3D way you can do so here.
First, here is the S100A10 protein in a “ball and stick” rendering, where each atom is a ball and the connections between atoms are the sticks:
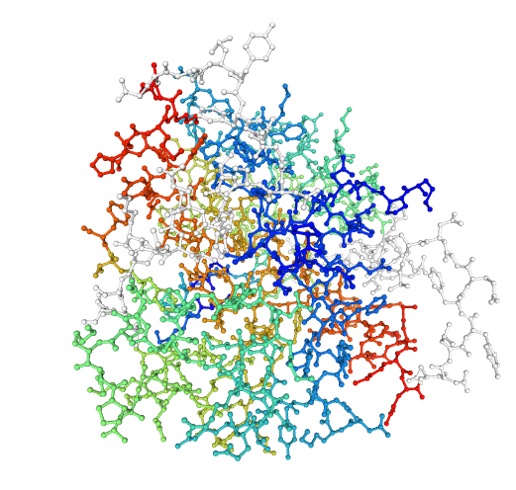
Here is a “spacefill” rendering:

Here is a “surface” rendering:
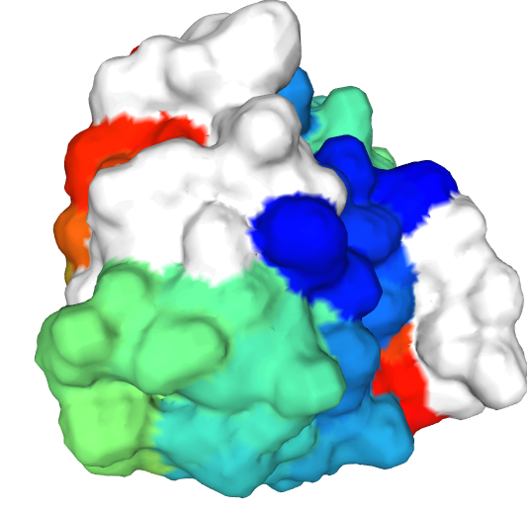
And finally, here is a cartoon rendering (the spirals are an abstract non-literal representation of a 3D motif called an “alpha helix” that is seen in many proteins):
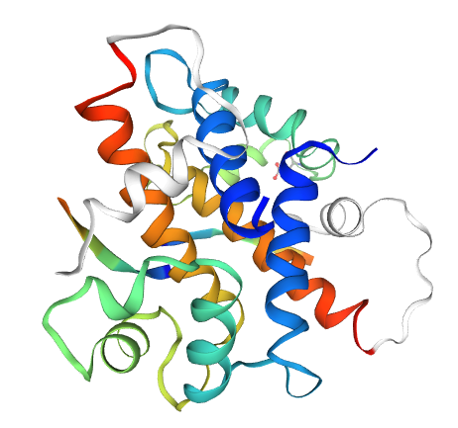
Proteomics
As you may have guessed, proteomics refers to the large-scale study of proteins. You may wonder why anyone would want to study proteomics if we have genomics (to understand all the instructions for building the body) and transcriptomics (to understand which machines are needed in which body parts).
Well, it turns out that transcriptomics isn’t a perfect representation of what machines are needed in which body parts, because RNA does not get made into proteins in a 1:1 ratio. In other words, you can’t just count up the number of RNA pieces made from Gene A and conclude that because there are 100 pieces of RNA from Gene A, that means that 100 versions of Protein A were produced. Sometimes, RNA is not made into protein at all. Sometimes, a certain piece of RNA is used to make many proteins, and other times, a piece of RNA is used to make only a few proteins. Therefore, proteomics provides a more direct way of assessing the total number of machines in a particular body part.
By combining the analysis of genomics, transcriptomics, and proteomics, we can gain a more complete understanding of a living creature’s health state and functionality.
The Inner Life of the Cell
To close I would highly recommend watching the YouTube video “The Inner Life of the Cell.” It depicts many of the processes we have talked about in this post, and is a stunning and biologically accurate depiction of the molecular machinery inside of all of us.
Summary
- DNA is the blueprint of life. The 3 billion letters of DNA in a human body contain all the instructions for building the body.
- Genomics is the study of DNA sequences on a large scale.
- Transcription is the process in which DNA is used to make RNA templates.
- Translation is the process in which RNA templates are used to make proteins.
- Transcriptomics is the study of RNA sequences on a large scale.
- Proteins are molecular machines that carry out different functions in the body.
- Proteomics is the study of proteins on a large scale.
About the Featured Image
The featured image is from Wikipedia, credit to William Crochot.
Want to be the first to hear about my articles bridging healthcare, artificial intelligence, and business—and get a free list of my favorite health AI resources? Sign up here.


Comments are closed.